- Publication
- May 19, 2022
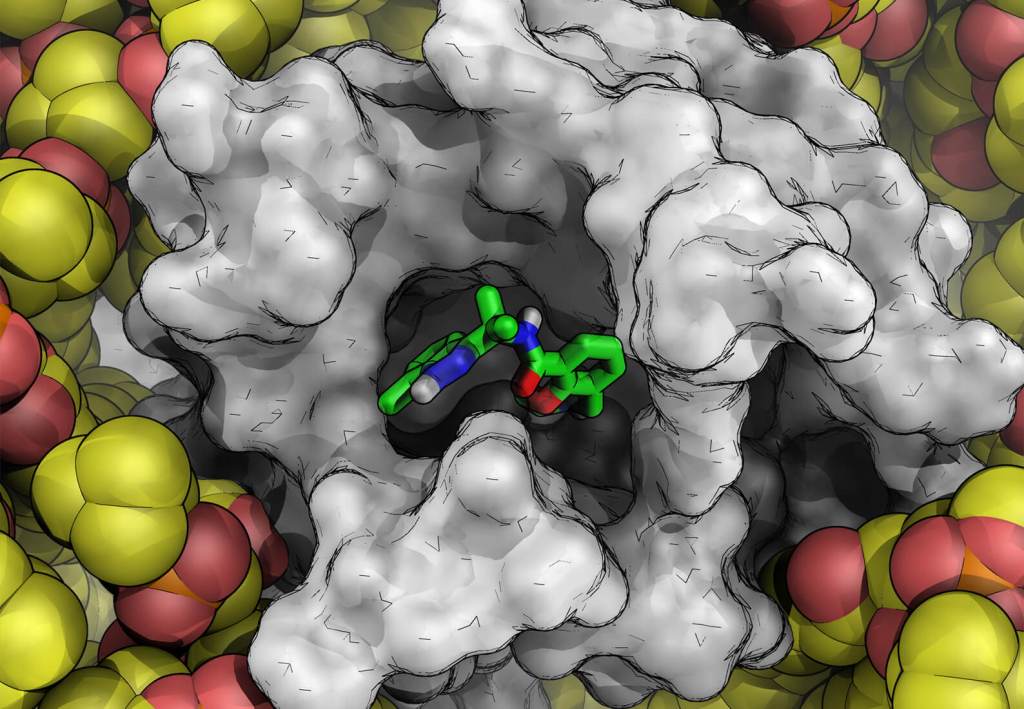
Plasticity in ligand recognition at somatostatin receptors
Michael J. Robertson, et al. Nature Structural & Molecular Biology, 2022, 29, 210-217- Publication
- May 19, 2022
Transferable Neural Network Potential Energy Surfaces for Closed-Shell Organic Molecules: Extension to Ions
Leif D. Jacobson, et al. JCTC, 2022, 18(4), 2354-2366- Publication
- May 19, 2022
AutoDesigner, a De Novo Design Algorithm for Rapidly Exploring Large Chemical Space for Lead Optimization: Application to the Design and Synthesis of D-Amino Acid Oxidase Inhibitors
Pieter H. Bos, et al. JCIM, 2022, 62(8), 1905-1915- Publication
- May 5, 2022
Induced-Fit Docking Enables Accurate Free Energy Perturbation Calculations in Homology Models
Tianchuan Xu, et al. Journal of Chemical Theory and Computation, 2022, 18(9), 5710-5724- Publication
- May 2, 2022
A Descriptor Set for Quantitative Structure-Property Relationship Prediction in Biologics
Kannan Sankar, et al. Mol Inform, 2022- Publication
- May 2, 2022
Accurate Prediction of Protein Thermodynamic Stability Changes upon Residue Mutation using Free Energy Perturbation
Guido Scarabelli, et al. JMB, 2022, 434(2)- Publication
- Apr 29, 2022
Intense bitterness of molecules: Machine learning for expediting drug discovery
Eitan Margulis, et al. Computational and Structural Biotechnology Journal, 2021, 19, 568-576- Publication
- Nov 2, 2021
The transcriptional corepressor CtBP2 serves as a metabolite sensor orchestrating hepatic glucose and lipid homeostasis
Sekiya, et al. Nat Commun, 2021, 12(1), 6315- Publication
- Oct 1, 2021
Biological activity validation of a computationally designed Rituximab/CD3 T cell engager targeting CD20+ cancers with multiple mechanisms of action
Cai, et al. Antib Ther, 2021, 4(4), 228-241- Publication
- Jul 6, 2021
Potency- and Selectivity-Enhancing Mutations of Conotoxins for Nicotinic Acetylcholine Receptors Can Be Predicted Using Accurate Free-Energy Calculations
Dana Katz, et al. Marine Drugs, 2021, 19(7), 367- Publication
- Jun 11, 2021
OPLS4: Improving Force Field Accuracy on Challenging Regimes of Chemical Space
Lu C.; Wu C.; Ghoreishi D.; Chen W.; Wang L.; Damm W.; Ross G.A.; Dahlgren M.K.; Russell E.; Von Bargen C.D.; Abel R.; Friesner R.A.; and Harder E.D., 2021
Latest insights from Extrapolations blog
Training & Resources
Online certification courses
Level up your skill set with hands-on, online molecular modeling courses. These self-paced courses cover a range of scientific topics and include access to Schrödinger software and support.
Free learning resources
Learn how to deploy the technology and best practices of Schrödinger software for your project success. Find training resources, tutorials, quick start guides, videos, and more.
Other Resources