FEP+ Protocol Builder
ML-powered optimization of FEP+ models for challenging targets

ML-powered optimization of FEP+ models for challenging targets
FEP+ Protocol Builder is an automated machine learning workflow for FEP+ model optimization. It saves scientists time and improves the chances of successfully enabling discovery with FEP+ by identifying an optimized predictive model for your target.
Fully automated FEP+ Protocol Builder generated optimized FEP+ protocol with improved pairwise RMSE compared to default model. All protocols were performed with the same 15 congeneric ligands with modifications at 2 R-groups with known affinity data from PDB ID: 2IVV. FEP+ Protocol Builder was applied to quickly and rigorously validate the amenability of the target to the Schrödinger platform.
| Workflow | Description | Library Size | Time to screen 4.0B (days) | Time to screen 6.5B (days) | Storage space for 6.5B (TB) |
|---|---|---|---|---|---|
| Quick Shape | Combination of 1D-SIM* prefilter and Shape CPU Screening | > 4.0 billion | 5.2 | 5.5‡ | 0.4 |
| Shape GPU | GPU-accelerated 3D screening | < 5.0 billion | 4.6 | 7.5‡ | 33 |
| Shape CPU | CPU-based 3D screening | < 10 million | NA | NA | NA |
In a recent study, FEP+ Protocol Builder outperformed human experts in producing a predictive model across ten diverse targets where default FEP+ settings did not produce an appropriately accurate protocol (RMSE > 2.5 kcal/mol) and were able to generate FEP+ models for systems where experts failed.
FEP+ Protocol Builder was run in a completely automated fashion following a rigorous training/test set split. On average, the reduction in turnaround time for the final optimization model went from 27 days to 7 days, a 4x acceleration, saving on average 20 days per project.
| Disease area | Target class | Target | Expert Protocol RMSE (kcal/mol) | FEP+ Protocol Builder RMSE (kcal/mol) |
|---|---|---|---|---|
| Oncology | Bcl-2 | MCL1 | 1.5 |
1.1 |
| Neurology | ATPase | P97 | 1.3 |
1.0 |
| Oncology | Nuclear receptor | ESR1 | 3.1 |
2.0 |
| Pain, addiction, oncology | GPCR | mOR | 2.4 |
2.2 |
| Pain, addiction | GPCR | dOR | 2.2 |
1.3 |
| Hematology, oncology | ADP-ribosyltransferase | TNKS2 | 2.2 |
1.1 |
| Pain, addiction, neurology | GPCR | KOR | 2.1 |
1.7 |
| Renal | Aspartic protease | Renin | 1.8 |
1.6 |
| Oncology and rheumatology | Cysteine protease | MALT1 | 2.5 |
1.5 |
| Oncology | Receptor tyrosine kinase | RET | 1.9 |
0.8 |

FEP+ is a powerful predictive technology in drug discovery – with applications from hit discovery through lead optimization. A critical first step in deploying FEP+ is to validate and optimize the model for the protein-ligand system of interest.
Watch Webinar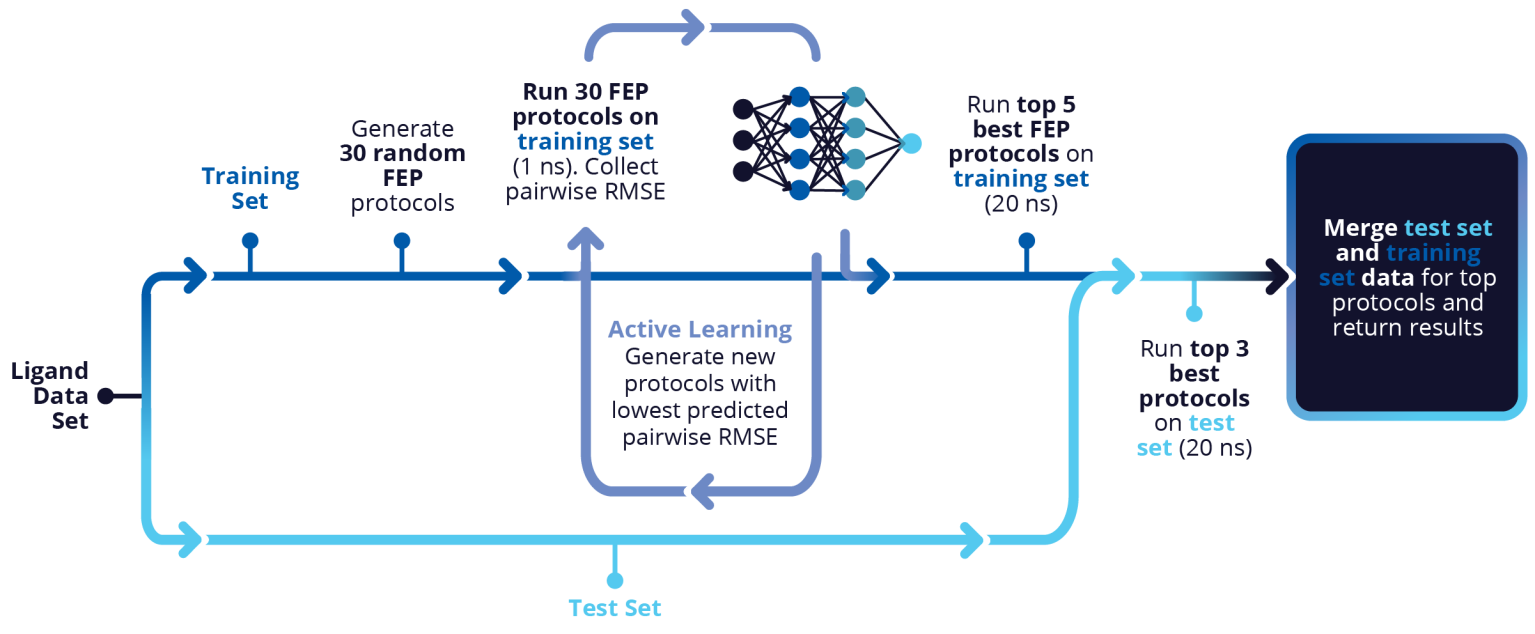
de Oliveira C, et al. J. Chem. Inf. Model. 2023, 63, 17, 5592–5603.
Abel R, et al. Acc. Chem. Res. 2017, 50, 7, 1625–1632.
Abel R, et al. J. Am. Chem. Soc. 2015, 137, 7, 2695–2703.