In this Stories from Drug Discovery series, Dr. Sayan Mondal, a Research Leader in Schrödinger’s Drug Discovery Group, demonstrates modeling strategies used from start to finish in a two-year oncology project in this short, 10-minute video. His team was tasked to first discover multiple potent, highly ligand-efficient lead series and then optimize them to find the potent compounds with balanced ADME profiles for development candidate nomination.
Stories from drug discovery: Modeling strategies in the pursuit of development candidate in oncology program 1

Summary
Dr. Sayan Mondal was the Modeling Lead on a recent drug discovery project at Schrödinger that was tasked with rapidly discovering a potentially best-in-class Type 1 kinase inhibitor in the backdrop of a competitor clinical compound entering Phase 2. The team’s goal was to have the key compounds for development candidate nomination in hand within two years from start to finish.
In this video, Dr. Mondal describes the modeling strategies the team used to efficiently overcome the typical challenges that often slow down programs like this or take them off the track. The team’s strategies delivered very potent, optimal compounds in multiple distinct chemical series as candidates for development candidate nomination within two years.
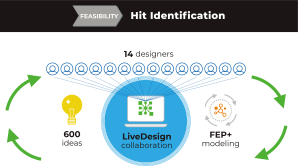
Feasibility Stage: Hit Identification
The project began with the feasibility stage and had two goals:
- Discover novel and potent chemical series
- Confirm that Schrödinger’s FEP+ is performing well for this program prospectively
Using LiveDesign as their centralized platform, the team crowd-sourced ideation in an iterative progression from the drug discovery group at large. Ideas were modeled with FEP+, and the FEP+ data were collated in LiveDesign to inspire new rounds of ideation. If an idea looked promising, the team would expand the model around it to come up with a final list of compounds.
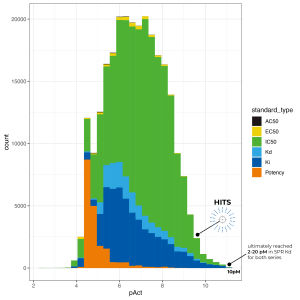
21 out of 23 predictions matched the experimental data with overall Mean Unsigned Error (MUE) well within one log order. The team identified five novel picomolar (pM) cores that were highly ligand efficient, ultimately reaching 2-20 pM in SPR binding assay with multiple compounds — among the most potent non-covalent kinase inhibitors known —for both the lead and the backup series.
Overcoming Selectivity, Permeability, and Solubility Challenges
Selectivity
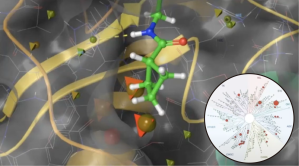
Selectivity challenges are common in kinase projects. Across both series, all tested compounds emerging from the feasibility stage were binding to a kinase with known cardiovascular toxicity, in addition to several other off target kinases. In order to dial out that liability, the team decided to use the new induced-fit docking engine, IFD-MD, to develop a putative binding pose for use in FEP+ off target modeling, instead of waiting for several months to solve a crystal structure of their series in the kinase off target.
Within days of receiving the kinase subpanel data, the team was able to get a reliable predictive FEP+ model for the key off-target that they were able to validate with compounds in the recent synthesis queue, where the predicted potency and experimental data were found to be in good agreement with each other.Thus, within two weeks the team was able to have a prospectively predictive model which they coupled with two design strategies for proof of concept (PoC) compounds for gaining kinase selectivity.
With this, the team was able to quickly find a PoC compound that was tolerated and maintained 100 pM potency on-target, but lost potency in the off-target kinase. In the DiscoverX kinase panel, the compound tested at 1µM (10,000x higher concentration than their on-target potency), and was quite clean, making it an excellent proof of concept for the model and the strategy.
Passive Membrane Permeability
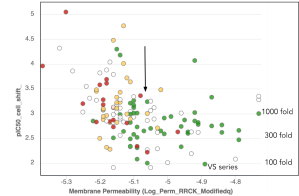
Passive Membrane Permeability RRCK is a fast method that can triage a large list of ideas before heavy modeling, reserving GPU resources for modeling only cell permeable chemical matter. RRCK modeling can inform project strategy in terms of whether and where more polarity can be tolerated in a series to achieve other ADME goals such as clearance and solubility.
In the beginning months of the program, the team was making compounds that were potent on-target in binding and biochemical assays, but were shifted in the cell potency by 3-5 log orders, a typical problem for kinase projects. After collecting rounds of data, the team found that the RRCK predictions could explain the experimental data with a thresholding effect, with an atypically aggressive cutoff of around -5.1 (cutoffs can be project and series dependent, tend to be around -5.5 in our experience). The team shifted their modeling strategy to meet the -5.1 cutoff, controlling the cell shift to 2-3 log orders.
Solubility
FEP+ solubility calculations helped transform the backup series from an insoluble to a soluble, potent, permeable regime within one round of synthesis prioritization, with an MUE of 0.5 logS vs. the experimental kinetic solubility in the program. This will be covered in detail in a future case study.
Conclusion
By implementing various modeling strategies in the Schrödinger platform throughout the lifecycle of the drug discovery program, the team was able to overcome challenges that are pervasive in kinase programs and get to pre-DC compounds within two years in distinct chemical series.
- Using FEP+ and RRCK permeability modeling, the team was able to obtain two distinct pM series that were both optimized to 2-20 pM potency and had good translation to cell potency
- Deploying selectivity FEP+ with an IFD-MD generated pose in a key kinase off target enabled the team to get to a rapid proof-of-concept compound without waiting for a crystal structure
- Working within LiveDesign allowed multiple contributors from across the globe to push the project ahead by testing, tracking and analyzing all relevant information in a centralized platform