APR 29, 2026
Using PyMOL to Visualize Protein Structure and Molecular Interactions in the Biology Classroom
Understanding three dimensional protein structure can be difficult for students when concepts are presented only through static textbook images. Visualization tools can improve spatial understanding and help students connect structure with biological function. This lightning talk demonstrates how PyMOL can be used as an accessible computational tool to enhance teaching of protein structure and molecular interactions using hemoglobin as a model system.
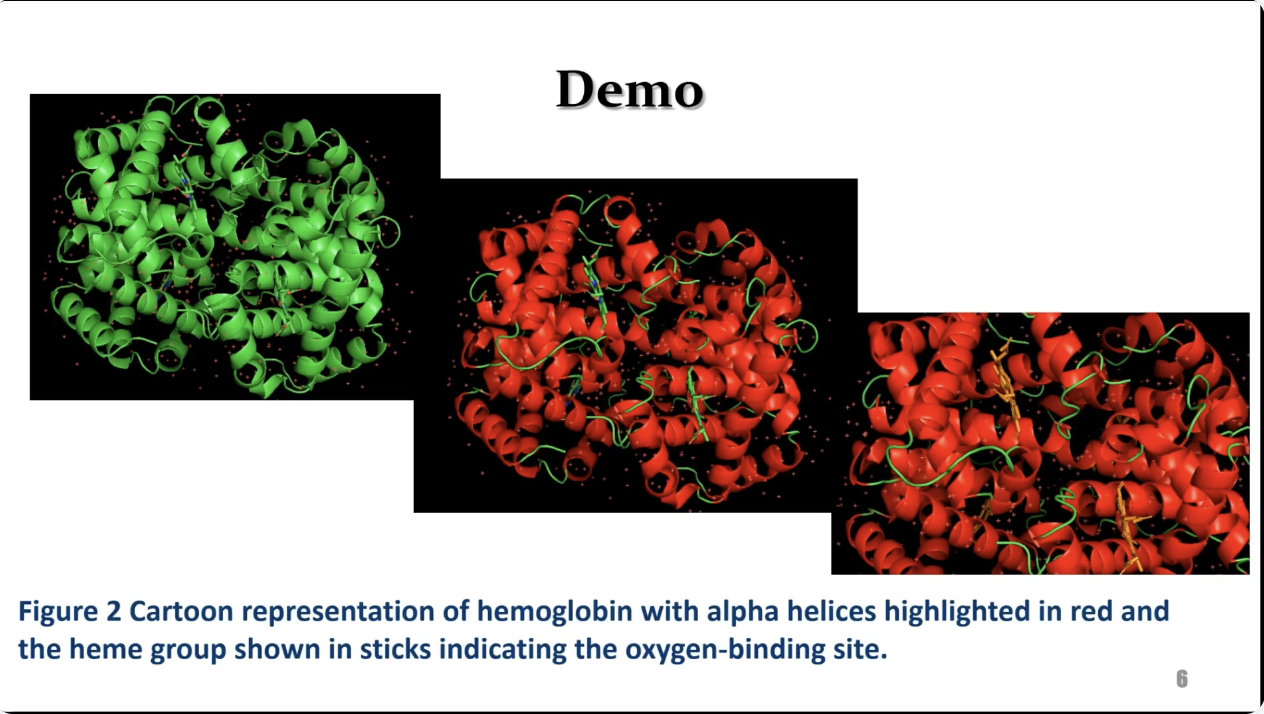
Hemoglobin is a well characterized protein with clear secondary and quaternary structural features and a biologically relevant oxygen binding mechanism. In this session, participants will see a guided demonstration of how to load a Protein Data Bank file into PyMOL, visualize alpha helices and beta sheets, highlight heme groups, and examine subunit organization. The workflow is designed for classroom use and requires minimal computational background.
The presentation will also outline strategies educators can use to guide students in exploring structure function relationships through interactive visualization. Participants will receive a concise classroom ready resource that includes a selected PDB identifier, basic PyMOL commands, and guided student questions for immediate implementation in undergraduate biology or biochemistry courses.
This session highlights how structural visualization can transform abstract molecular concepts into engaging learning experiences.
Our Speaker

Aruna Arumugam Chockalingam
Research Scholar, Asia University