- Publication
- May 8, 2026
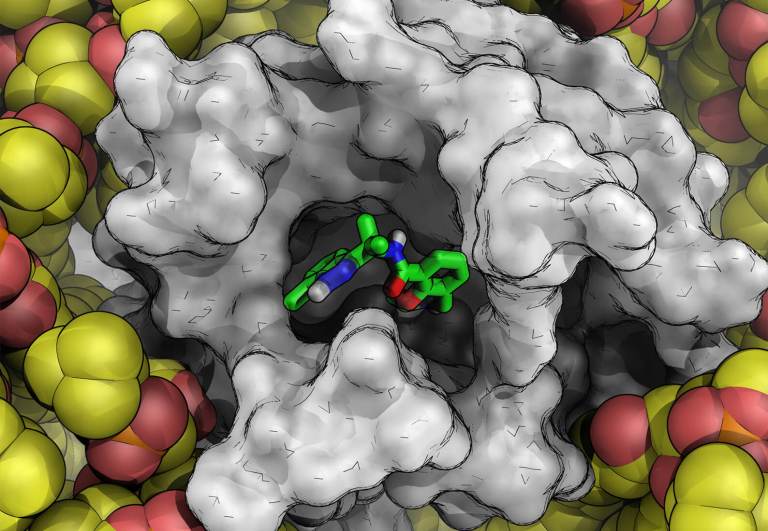
Discovery of 2H-Pyrrolo[3,4-c]pyridin-3-one Derivatives as Type-III c-MET Inhibitors Enabled by Free-Energy Perturbation CalculationsCl
Therrien, et al. ACS Medicinal Chemistry Letters, 2026
- Publication
- Apr 8, 2026
Structure-Based Discovery of Imidazo[4,5-c]pyridine SARM1 Modulators Showing Paradoxical Activation
Albanese, et al. Journal of Medicinal Chemistry, 2026, 69(8), 9521–9536
- Publication
- Mar 21, 2026
Structure-Based Calculation of Excipient Effects on the Viscosity of Concentrated Antibody Solutions
Shelley, et al. mAbs, 2026, 18(1)
- Publication
- Feb 16, 2026
Accurate physics-based flexible docking of macrocyclic ligands
J, et al. Journal of Medicinal Chemistry, 2026, 69(4), 4745–4754
- Publication
- Jan 16, 2026
Prediction of Atropisomerism for Drug-like Molecules
Balduf, et al. Journal of Chemical Information and Modeling, 2026, 66(3), 1675–1687
- Publication
- Dec 12, 2025
Unique Interactions of Novel Rufomycin ‘Click-Chemistry’ Analogs with Mtb ClpC1 and Implications
Ratia, et al. Journal of Medicinal Chemistry, 2025, 68(24), 26298–26310
- Publication
- Dec 3, 2025
Glide WS: Methodology and Initial Assessment ofPerformance for Docking Accuracy and Virtual Screening
Friesner, et al. Journal of Chemical Theory and Computation, 2025, 21(24), 12696–12708
- Publication
- Dec 3, 2025
Glide WS: Methodology and Initial Assessment of Performance for Docking Accuracy and Virtual Screening
Friesner, et al. Journal of Chemical Theory and Computation, 2025, 21(24), 12696–12708
- Publication
- Dec 1, 2025
Rapid estimation of protein folding pathways from sequence alone using AlphaFold2
Chang, et al. Nature Communications, 2025, 17, 170
- Publication
- Oct 13, 2025
Accelerated in silico discovery of SGR-1505: A potent MALT1 allosteric inhibitor for the treatment of mature B-cell malignancies
Nie Z, et al. J. Med. Chem., 2025
- Publication
- Oct 12, 2025
Discovery of highly potent noncovalent inhibitors of SARS-CoV-2 main protease through computer-aided drug design
Okabe A, et al. J Med Chem, 2025
- Publication
- Sep 17, 2025
Accurate hydration free energy calculations for diverse organic molecules with a machine learning force field
Xie, et al. ChemRxiv, 2025, 1, Preprint
Events
 Webinar
Materials Science
Webinar
Materials Science
- May 29, 2026
Beyond the bench: Getting started with molecular dynamics simulations
Join Schrödinger’s Katie Dahlquist, as she’ll show you how Desmond can be used to improve your development.
 Webinar
Materials Science
Webinar
Materials Science
- Jun 2, 2026
Accessible and automated computational catalyst discovery and reactivity optimization
In this webinar, we will demonstrate how an end-user physics–AI platform removes barriers to entry, making this process accessible to both experts and non-experts while enabling seamless scalability.
 Event
Life Science
Event
Life Science
- Jun 2nd-4th, 2026
Computational and Medicinal Chemistry by the Lake 2026
Schrödinger is excited to be participating in the Computational and Medicinal Chemistry by the Lake 2026 conference taking place on June 2nd – 4th in Kuopio, Finland.
Webinars
 Webinar
Life Science
Webinar
Life Science
- Jun 3, 2026
Biologics modeling for wet lab scientists: Detecting and deprioritizing dead ends before they reach the bench
Join us to learn how to detect and deprioritize high-risk candidates, effectively discarding developability dead-ends before they ever reach the bench.
 Webinar
Life Science
Webinar
Life Science
- Jun 4th-11th, 2026
Generative Glide: AI-driven ultra-large virtual screening for real-world drug discovery
Join us as we go beyond slides and run a demo of the workflow, showing how Generative Glide performs in practice from setup through results.
 Webinar
Life Science
Webinar
Life Science
- Jun 10th-16th, 2026
Expanding RetroSynth: Dynamic AI-driven synthesis planning proven to move your designs from bench to synthesis
In this webinar, we revisit RetroSynth, Schrödinger’s AI-powered solution for intelligent synthesis planning.
Documentation
- Documentation
Learning Path: Oligonucleotide Modeling
A structured overview of tools and workflows for nucleic acids in drug discovery.
- Documentation
WaterMap
Efficiently converged MD simulations are run with explicit water molecules, and resultant trajectories are analyzed to cluster hydration sites.
- Documentation
SiteMap
Identify binding sites, including allosteric binding sites and protein-protein interfaces, and evaluate their druggability.
Tutorials
- Tutorial
Structure-Based Virtual Screening using Glide
Prepare receptor grids for docking, dock molecules and examine the docked poses.
- Tutorial
Ligand Binding Pose Generation for FEP+
Generate starting poses for FEP simulations for a series of BACE1 inhibitors using core constrained docking.
- Tutorial
Homology Modeling of Protein-Ligand Binding Sites with IFD-MD
Create a homology model of TYK2 from JAK3 and including a bound ligand. Compare this model with the crystal structure for TYK2 bound to 4GIH.
Training Videos
 Video
Life Science
Video
Life Science
Getting Going with Maestro BioLuminate
A free video series introducing the basics of using Maestro Bioluminate.
 Video
Life Science
Video
Life Science
- Video
Introducing Ligand Designer
An overview of the LigandDesigner workflow, Editing in 2D and 3D, using display options and overlays, and accessing the Admin Panel.
Publications
- Publication
- May 8, 2026
Discovery of 2H-Pyrrolo[3,4-c]pyridin-3-one Derivatives as Type-III c-MET Inhibitors Enabled by Free-Energy Perturbation CalculationsCl
Therrien, et al. ACS Medicinal Chemistry Letters, 2026
- Publication
- Apr 8, 2026
Structure-Based Discovery of Imidazo[4,5-c]pyridine SARM1 Modulators Showing Paradoxical Activation
Albanese, et al. Journal of Medicinal Chemistry, 2026, 69(8), 9521–9536
- Publication
- Mar 21, 2026
Structure-Based Calculation of Excipient Effects on the Viscosity of Concentrated Antibody Solutions
Shelley, et al. mAbs, 2026, 18(1)
Case Studies
 Case Study
Life Science
Materials Science
Case Study
Life Science
Materials Science
 Case Study
Life Science
Case Study
Life Science
 Case Study
Life Science
Case Study
Life Science
White Papers
 White Paper
Life Science
White Paper
Life Science
- Jan 29, 2026
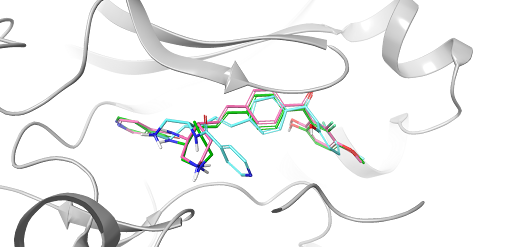
FEP+ Pose Builder — maximizing utility and productivity in FEP simulations
FEP+ Pose Builder is a methodological advancement introduced as an integrated feature to drastically enhance accessibility, user-friendliness, and productivity within the FEP+ pipeline.
 White Paper
Life Science
White Paper
Life Science
- Oct 29, 2024
20 Years of Glide: A Legacy of Docking Innovation and the Next Frontier with Glide WS
Glide has long set the gold standard for commercial molecular docking software due to its robust performance in both binding mode prediction and empirical scoring tasks, ease of use, and tight integration with Schrödinger’s Maestro interface and molecular discovery workflows.
 White Paper
Life Science
White Paper
Life Science
Quick Reference Sheets
- Quick Reference Sheet
Force Field Builder
A one-page guide to calculate missing torsion parameters for ligands using the Force Field Builder panel.
- Quick Reference Sheet
Ligand Interaction Diagram
A one-page guide to using the Ligand Interaction Diagram for examining ligand-receptor interactions.
- Quick Reference Sheet
GlideMap
A one-page guide to using the GlideMap GUI for ligand placement guided by experimental density.

Latest insights from Extrapolations blog
Training & Resources
Online certification courses
Level up your skill set with hands-on, online molecular modeling courses. These self-paced courses cover a range of scientific topics and include access to Schrödinger software and support.
Free learning resources
Learn how to deploy the technology and best practices of Schrödinger software for your project success. Find training resources, tutorials, quick start guides, videos, and more.