Release Notes
Release 2026-1

Small Molecule Drug Discovery
Platform Environment
Maestro Graphical Interface
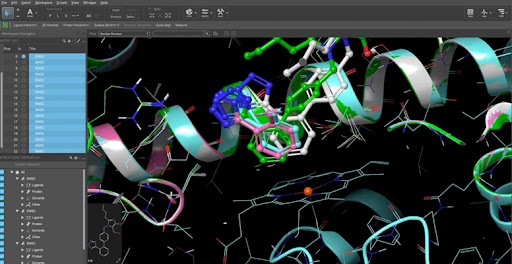
- Redesigned Surface Manager – Control complex visualizations effortlessly with a modern, persistent interface that allows for real-time, non-modal editing of surface styles, colors, and transparency
- Persistent measurements – Geometric measurements now persist with the entries, allowing for uninterrupted structural comparison across multiple conformers and states
- Standalone density map import – Accelerate Cryo-EM and crystallography workflows with direct, standalone density map import
- Maestro Assistant modes (open beta) – A context-aware AI partner that intelligently toggles between ‘Ask’, ‘Execute’, and ‘Auto’ modes to seamlessly bridge documentation and direct action
Binding Site & Structure Analysis
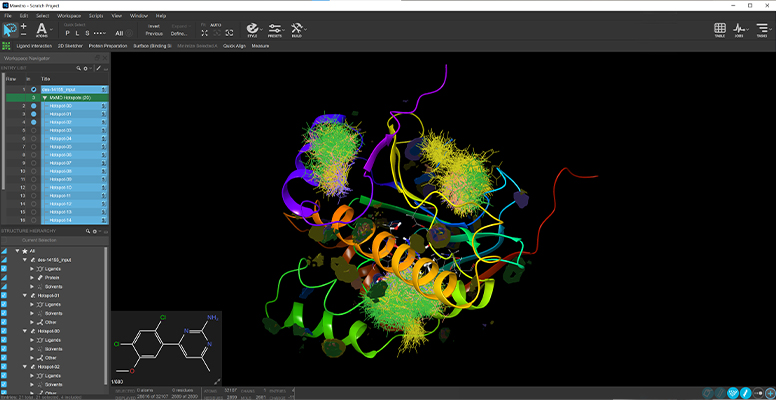
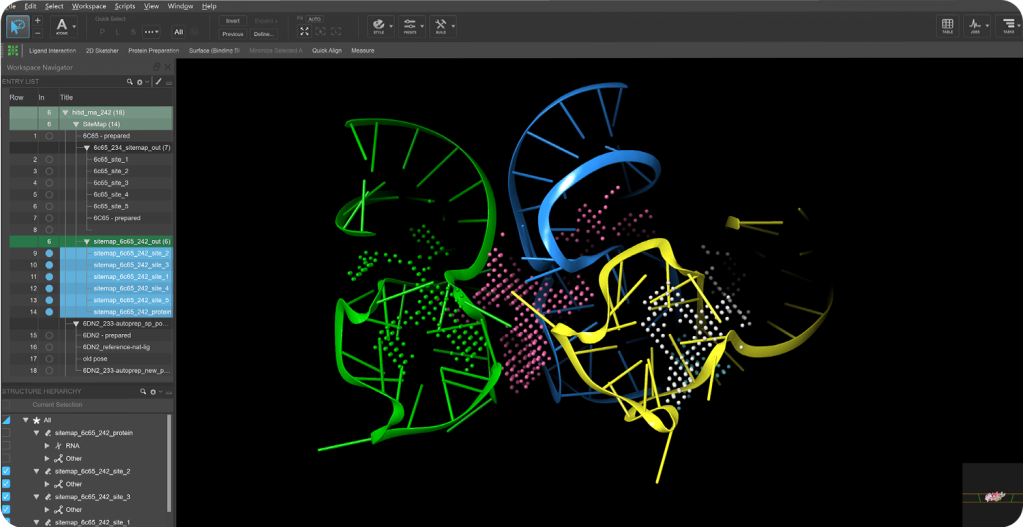
Mixed Solvent MD (MxMD)
- Added support to command line MxMD driver to seamlessly execute all simulations and compile results from combined MxMD/SiteMap cryptic pocket identification workflow
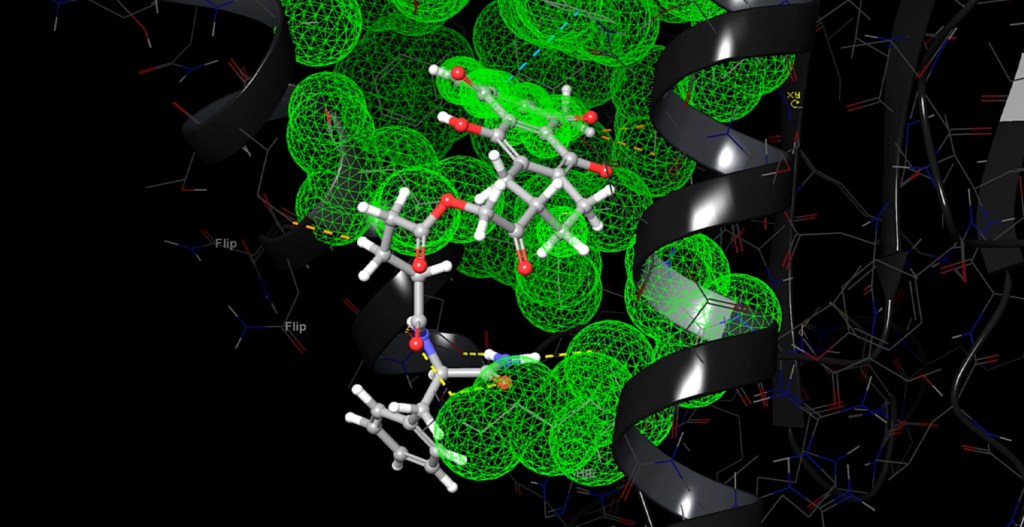
WaterMap
- WaterMap now supports use of the OPLS_2005 forcefield and TIP4P water model
Hit Identification & Virtual Screening
Docking
- Understand, optimize, and troubleshoot native redocking experiments with new Docking Report to maximize docking performance
Lead Optimization
FEP+
- GraphDB/Web services can optionally download only the primary FMP file and not the FMPdb reducing time to analyzing results
- FEP+ workflows now support execution on cost-effective preemptible nodes
- Use 2D Sketcher to define Core SMARTS for FEP+
- Improved handling of categorical assay data in FEP+ statistical analysis
FEP+ Protocol Builder
- Sample three levels of salt concentrations in protocol optimization
- Sample automatic membrane placement in protocol optimization
FEP+ Pose Builder
- Automatically create accurate and clash-aware FEP-ready poses with FEP+ Pose Builder: Generate high-quality ligand alignments faster to run FEP+ at scale with an automated workflow designed for unbiased selection and robust atom-mapping
- LiveDesign FEP+ Pose Builder protocol now supports generating FMP files for cycle-closure FEP calculations
Quantum Mechanics
- Employ xTB and MLFFs including QRNN, MPNICE, and UMA in AutoTS from command line for shorter calculation times
- Faster batch calculations by optimized CPU core assignments for all multithreaded Jaguar batch calculations
Spectroscopy
- New corrections for C-Br and C-I bonds in 13C NMR spectra
De Novo Design
AutoDesigner – R-group Design
- Explore large chemical spaces to identify optimal R-groups with new AutoDesigner R-group Design Panel now accessible by setting a feature flag
- Added optional QED score (Quantitative Estimate of Drug-Likeness) for AutoDesigner R-group Design ideas
AutoDesigner – Core Design
- Explore large chemical spaces to identify optimal core replacements with new AutoDesigner Core Design Panel now accessible by setting a feature flag
- Added optional QED score (Quantitative Estimate of Drug-Likeness) for AutoDesigner Core Design ideas
Education Content
- Introducing a new, more interactive tutorial experience, letting users intuitively click through the steps of a workflow right in the tutorial. This approach is being piloted in two tutorials.
- Updated Tutorial: Structure-Based Virtual Screening Using Glide
- Updated Tutorial: Introduction to Structure Preparation and Visualization
- Fully re-worked learning content on Antibody Modeling:
The “Antibody Visualization and Modeling in BioLuminate” tutorial has been replaced by a new suite of tutorials organized in a learning path together with other helpful resources.- New Learning Path: Antibody Modeling
- New Tutorial: Antibody Structure Prediction and Visualization with BioLuminate
- New Tutorial: Humanizing Antibody Structures with BioLuminate
- New Tutorial: Antibody – Antigen Docking with PIPER
- New Tutorial: Improving Antibody Stability/Affinity Using MM-GBSA Residue Scanning
- New Tutorial: Generating ternary complex structures to enable rational design of targeted protein degraders
- Updated Tutorial: BACE1 Inhibitor Design Using Free Energy Perturbation
- New Quick Reference Sheet: Force Field Builder
- New Quick Reference Sheet: Ligand Interaction Diagram
Biologics Drug Discovery
- Release of MacroMolecular Pose Filter – Select the most plausible or relevant structural models of a macromolecular complex from a larger set of generated possibilities
- New protein descriptor – ASPmax – Added ASPmax (Maximum Average Surface Property) to our descriptor set. Used to predict the retention time of proteins in hydrophobic interaction chromatography (HIC) columns and aggregation risk
- Search and filter non-standard residues – Support for text-based searching and filtering of non-standard residues based on labels in the Name, Code, and Description columns
- Residue Lookup in the MMGBSA residue scanning panel – Quickly search and find residues to mutate
- Classification of residue scanning results – Color codes residue scanning results to designate positive, neutral or negative mutational variants based on energy score cut-offs
Materials Science

GUI for Quantum ESPRESSO
Product: Quantum ESPRESSO (QE) Interface
- Defect Formation Energy: Workflow solution to analyze point defects in crystals
- Options to set frequency cutoff / harmonic threshold in the Phonon DOS Viewer
- Support for TB09 density functional (command line)
- Support for rVV10-SCAN density functional (command line)
- Support for NpT ensemble in QE BOMD simulations (command line)
- (+MATSCI_NEB_MLFF) MLFF integration in NEB
MS Surface
Product: MS SurfChem
- Desorption Enumeration: WAM to open results in Adsorption Energy
Microkinetics
Product: MS Microkinetics
- Option to view selectivities and degrees of selectivity control
- Option to load/save archived MKM output
- Option to export reaction view as an image (PNG) file
- Results from individual stages made visible from the analysis panel
Optoelectronics Genetic Optimization
Product: Genetic Optimization (GA)
- Support for setting target property based on models from ML Property Prediction
Active Learning Optoelectronics
Product: Active Learning Optoelectronics
- Option to set target values excluded from optimizations
Reactivity
Product: MS Reactivity
- Nanoreactor: Option to specify separate hosts for driver and subjobs
- Nanoreactor: (+NANOREACTOR_AUTOTS) Automatic transition state search for elementary reaction network calculations via AutoTS
- Nanoreactor: Option to skip generating trajectory files
- Nanoreactor: User control over the time interval between trajectory frames
- Reaction Network Profiler: Option to refine conformer geometries using UMA (MLFF)
- Reaction Network Profiler: Option to assign stoichiometric multipliers for reactants
Transport Calculations via MD simulations
Product: MS Transport
- Ionic Conductivity: Support for MLFF
Dielectric properties
Product: MS Dielectric
- Complex Permittivity: Support for multi-component systems
Coarse-Grained (CG) Molecular Dynamics
Product: MS CG
- CG FF Builder: Improved detection and mapping of non-isomorphic residues
- CG FF Builder: Automated particle naming scheme with chemical context
- CG FF Assignment: Up to 15x speed-up for models with a large number of particle types
- Coarse-Grained Mapping: Residue number and name retained through mapping
- Coarse-Grained Mapping: Option to import SMARTS patterns from previous use
- Speed-up for DPD simulations of up to 30% with improved cutoff margins
Complex Bilayer Builder
Product: MS Complex Bilayer Builder
- Complex Bilayer: (+COMPLEX_BILAYER_BUILDER_EXTENDED_LIPID_LIB) Expanded list of default lipids
- Complex Bilayer: Increased limit for water padding depth to 5000 Å
- Complex Bilayer: Support for custom-trained OPLS
- Membrane Analysis: (+MEMBRANE_ANALYSIS_PREP_FOR_FEP) Support for generating poseviewer formatted files compatible with FEP calculations
- Membrane Analysis: Support for applying multiple leaflet-finding algorithms
Materials Informatics
Product: MS Informatics
- Machine Learning Property: Improved panel interface for model selection
- MD Descriptors: User control over simulation system size (max # of atoms)
- MD Descriptors: Support for formulation input with path assigned to structure files
- MLFF Calculations: Option to set constraints to atomic positions
- MLFF Calculations: Support for running on GPU nodes
Formulation ML
Product: MS Formulation ML
- Formulation ML: ‘Learned Fingerprint’ as a new option to feature space
- Formulation ML: Advanced option to process (‘impute’) training set data with partially missing descriptors
- Formulation ML: Improved UI for parity plot
- Formulation ML: Support for building machine learning models using training datasets with missing chemical (SMILES) information
- Formulation ML: Target property displayed in the ‘Performance’ tab
- Formulation ML: User control over correlation threshold between features
- Formulation ML Optimization: Support for custom-ingredient descriptors
- Formulation ML Optimization: Support for multi-CPU parallelization
- Formulation ML Optimization: Option to use genetic algorithms for formulation optimization
Layered Device ML
Product: MS Layered Device ML
- OLED Device ML: User control over correlation threshold between features
- OLED Device ML: Target property displayed in the ‘Performance’ tab
MS Maestro Builders and Tools
- Adsorption Enumeration: WAM to open results in Adsorption Site Finder / Adsorption Energy
- Adsorption Site Finder: WAM to open results in Adsorption Energy
- Adsorption Enumeration: Improved organization of output structures in the Project Table
- Adsorption Site Finder: Up to 200x of speed-up for jobs using MLFF
- Clean Up Structures: Support for MLFF
- Disordered System: Option to define residue name for components
- Support for converting *.vis files to *.cub formatted files (command line)
- Polymer: Import of coupling probabilities from a CSV formatted file
- Polysaccharide: (+POLYSACCHARIDE_BUILDER) Simplified model building solution for linear-chain polysaccharides
- Single Complex: Updated list of bridging ligands
- Query Bonds: Display of polyhedra for molecular crystals
- Query Bonds: Search for and modification of non-bonded atom pairs
Classical Mechanics
- Droplet: Support for using pre-assembled droplet models
- Droplet: Support for computing contact angles with hydrate surfaces
- Elastic Constants: Support for MLFF
- (+ALLOW_OLD_FORCEFIELD_PARAMETERS) Support for running MD using OPLS4/OPLS5 parameters with backwards compatibility (2025-4 and older)
- MD Multistage: FF type for the input structure displayed in the panel
- Stress Strain: Support for MLFF
- Stress Strain: Option to use velocities from previous strain steps
- Surface Tension: Improved analysis with block averaging scheme
- Tg: (+THERMOPHYSICAL_PROPERTIES_MLFF) Support for MLFF
- Umbrella Sampling: Workflow solution to run umbrella sampling algorithm for small molecules near lipid and surfactant bilayers
- Umbrella Sampling: Displaying quantity of overlap between windows
- Viscosity: Adjusted default timestep (0.5 fs) for MLFF simulations
Quantum Mechanics
- Adsorption Energy: Option to pre-optimize structures with MLFF
- Adsorption Energy: Support for atomic positional constraints with MLFF
- Crest: (+MATSCI_CREST_QCG) CREST Quantum Cluster Growth Utility
- QM Multistage: Option to select GFN2-xTB from the list of theory
- Optoelectronic Film Properties: Display of refractive index ratio per molecular species
- Reaction Network Viewer: Comprehensive analysis viewer GUI for viewing networks created by Reaction Network Profiler and Nanoreactor
Education Content
- New Tutorial: Catalytic Selectivity Through Microkinetic Modeling
- Updated Tutorial: Defect Formation Energy Calculation
- Updated Tutorial: Nanoreactor
- Quick Reference Sheet: Clean Up Structures
Education Content
Life Science
- Introducing a new, more interactive tutorial experience, letting users intuitively click through the steps of a workflow right in the tutorial. This approach is being piloted in two tutorials.
- Updated Tutorial: Structure-Based Virtual Screening Using Glide
- Updated Tutorial: Introduction to Structure Preparation and Visualization
- Fully re-worked learning content on Antibody Modeling:
The “Antibody Visualization and Modeling in BioLuminate” tutorial has been replaced by a new suite of tutorials organized in a learning path together with other helpful resources.- New Learning Path: Antibody Modeling
- New Tutorial: Antibody Structure Prediction and Visualization with BioLuminate
- New Tutorial: Humanizing Antibody Structures with BioLuminate
- New Tutorial: Antibody – Antigen Docking with PIPER
- New Tutorial: Improving Antibody Stability/Affinity Using MM-GBSA Residue Scanning
- New Tutorial: Generating ternary complex structures to enable rational design of targeted protein degraders
- Updated Tutorial: BACE1 Inhibitor Design Using Free Energy Perturbation
- New Quick Reference Sheet: Force Field Builder
- New Quick Reference Sheet: Ligand Interaction Diagram
Materials Science
- New Tutorial: Catalytic Selectivity Through Microkinetic Modeling
- Updated Tutorial: Defect Formation Energy Calculation
- Updated Tutorial: Nanoreactor
- Quick Reference Sheet: Clean Up Structures
LiveDesign
What’s New in 2026-1
- Biologics
- Design new biologics with point mutations using natural monomers
- Upload custom monomers via API access, and view the monomers in the sequence viewer
- Search for a subsequence within an annotated region of a Biologics entity, by selecting the annotation and numbering scheme from pre-filled dropdowns
- Users can now color residues by property in the sequence viewer with any model that outputs the per-residue property/properties and color scheme mapping(optional) in the specified format, or with any Freeform column that contains the output
- The “Biologics” and “Generic Entity” options in the “Type” dropdown menu of the Advanced search panel, have been unified to “Biologics/Others”
- View branched and cyclic peptides in the sequence viewer
- LiveDesign AI Assistant: interact with LiveDesign using an AI assistant to create Freeform and Formula columns, create and update coloring rules, perform data analyses and plot data within the Assistant, and instantly find help documentation. Note that this capability requires the LiveDesign ML plugin
- Project Dashboards
- View a project activity stream of newly added assay data and comments using the new “Activity” section
- Entity count statistics and recently added entities shown in the Project Dashboard will now include all entities that are searchable to the project (e.g., compounds imported to unrestricted projects), instead of entities specifically imported to the project
- Models: Column as parameter models that use a 3D column as input now have access to the favorited pose, and the pose order that is represented in the LiveReport
- UX Improvements
- Filter out un-run, failed, and pending model cells from your LiveReport
- Drag a compound structure from the main spreadsheet directly to the Design, Search,or Advanced search panels without opening the panel first
- Close LiveReport tabs by clicking on them with a middle mouse button click
- 3D Visualizer: apply Stereolabels, Element label, and Atom Number labels to the selected atoms/residues/Chains
What’s Been Fixed
- Formulas that used the Lot Registration Date column as input would fail to calculate, and cause a red error bar to appear on the LiveReport. Those formulas now calculate correctly and do not cause an error
- Icons within the main spreadsheet cells to view a pose in the 3D visualizer, or in Maestro, would disappear when the cell was resized, and now remain visible
- Adding a click-to-run model column to a matrix widget would cause the widget to crash and not show data, and now click-to-run models appear correctly in the matrix widget
- Formulas using if() statements would fail to calculate when the if() statement used experimental assay cells that contained multiple values, in which one of those values was Null. Those formulas now calculate correctly
- The “Creating New Layout” dialog did not show an option to copy an existing layout, and now correctly shows that option.
- Editing a LiveReport’s title using the “Edit LiveReport Dialog” would result in moving the LiveReport to the Project Home folder, and now editing a LiveReport with that dialog will not move the LiveReport to a different folder
- Formatting option buttons on the Configure Matrix Widget dialog overlapped, which prevented accessing some formatting options, and now the buttons do not overlap
- Experimental assay data would show dashed lines underneath experimental values within the spreadsheet cells, and now do not show dashed lines
- The aggregateMax() and aggregateMin() formula functions now work for date columns
- Multi-chain or branched peptides are now accurately identified with incremental peptide numbers in their identifiers in sequence viewer.
- The sequence viewer now shows a message when TCRs with unsupported numbering schemes are used, and suggests to move to a supported numbering scheme for TCRs in the sequence viewer.
- When 3D visualizer is drilling down from the sequence viewer, selecting another entity in the sequence viewer does not reset the existing residue selection in the sequence viewer and 3D visualizer
- Parameterized models that used another model’s image columns as input would not calculate results, and now correctly calculate
- Recalculating a deleted model return would result in a permanently flashing cell, and now the cell will correctly show a Failed message
- When models returned a 3D output of type “other”, that 3D output was not accessible to other parameterized models, and now is accessible
- LDClient: the method get_models_by_name now includes an option to ignore archived models
Training & Resources
Online Certification Courses
Level up your skill set with hands-on, online molecular modeling courses. These self-paced courses cover a range of scientific topics and include access to Schrödinger software and support.
Tutorials
Learn how to deploy the technology and best practices of Schrödinger software for your project success. Find training resources, tutorials, quick start guides, videos, and more.