- Documentation
Learning Path: Oligonucleotide Modeling
A structured overview of tools and workflows for nucleic acids in drug discovery.
- Documentation
WaterMap
Efficiently converged MD simulations are run with explicit water molecules, and resultant trajectories are analyzed to cluster hydration sites.
- Documentation
SiteMap
Identify binding sites, including allosteric binding sites and protein-protein interfaces, and evaluate their druggability.
- Documentation
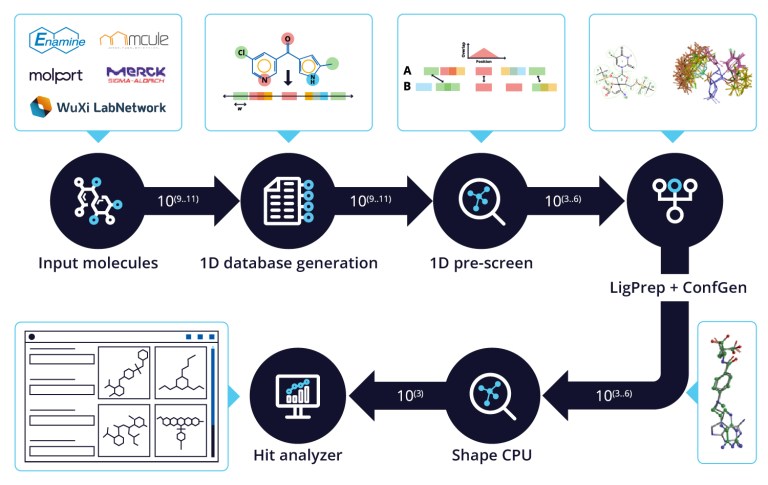
Shape Screening
A ligand-based workflow for efficiently screening ultra-large purchasable or synthesizable compound libraries.
- Documentation
QSite
A multi-scale simulation tool that utilizes the QM/MM method, which combines the principles of quantum mechanics and molecular mechanics.
- Documentation
QikProp
An advanced tool for predicting pharmacokinetic and physicochemical (ADME) properties of small organic molecules based on the full 3D molecular structure.
- Documentation
PrimeX
An advanced tool to facilitate protein crystal structure refinement and produces accurate structures more compatible with computational chemistry applications than those produced by other protein refinement programs.
- Documentation
Prime
A fully-integrated protein structure prediction solution that incorporates homology modeling and fold recognition into a single solution.
- Documentation
Phase
An intuitive pharmacophore modeling tool that allows assessment of compounds based on the steric and electronic features of molecules known to have biological activity.
- Documentation
OPLS4 and OPLS5 Force Field
A force field that is a model of the potential energy of a chemical system – a set of functions and parameters used to model the potential energy of the system, and thereby to calculate the forces on each particle.
- Documentation
Membrane Permeability
Calculate the passive membrane permeability of a set of congeneric ligands.
Events
 Event
Materials Science
Event
Materials Science
- Mar 31st – Apr 3rd, 2026
ICDT 2026
Schrödinger is excited to be participating in the International Conference on Display Technology, ICDT 2026 taking place on March 31st – April 3rd in Chongqing, China.
 Webinar
Life Science
Webinar
Life Science
- Apr 8, 2026
The unified antibody workbench: Optimizing antibody candidate selection through centralized team collaboration
Join us for an interactive webinar where we showcase the power of LiveDesign for Biologics in a live demo and walk you through how you can quickly start benefiting from this powerful enterprise informatics platform today!
 Webinar
Materials Science
Webinar
Materials Science
- Apr 9, 2026
Beyond the bench: Getting started with molecular dynamics simulations
Join Schrödinger’s Katie Dahlquist, as she’ll show you how Desmond can be used to improve your development.
Webinars
 Webinar
Life Science
Webinar
Life Science
- Apr 8, 2026
The unified antibody workbench: Optimizing antibody candidate selection through centralized team collaboration
Join us for an interactive webinar where we showcase the power of LiveDesign for Biologics in a live demo and walk you through how you can quickly start benefiting from this powerful enterprise informatics platform today!
 Webinar
Life Science
Webinar
Life Science
- Apr 16, 2026
Amplifying medicinal chemist impact with large-scale ideation, FEP+, machine learning, and retrosynthesis through LiveDesign
Join us to see how Schrödinger’s Enterprise Informatics Platform, LiveDesign, serves as the single terminal to bridge this gap.
 Webinar
Life Science
Webinar
Life Science
- Apr 23, 2026
Standing out in a competitive landscape: The power of structure-based biologics design
Join our upcoming webinar to learn how to leverage advanced in silico methods to de-risk your molecules and increase your experimental success rates.
Documentation
- Documentation
Learning Path: Oligonucleotide Modeling
A structured overview of tools and workflows for nucleic acids in drug discovery.
- Documentation
WaterMap
Efficiently converged MD simulations are run with explicit water molecules, and resultant trajectories are analyzed to cluster hydration sites.
- Documentation
SiteMap
Identify binding sites, including allosteric binding sites and protein-protein interfaces, and evaluate their druggability.
Tutorials
- Tutorial
Structure-Based Virtual Screening using Glide
Prepare receptor grids for docking, dock molecules and examine the docked poses.
- Tutorial
Ligand Binding Pose Generation for FEP+
Generate starting poses for FEP simulations for a series of BACE1 inhibitors using core constrained docking.
- Tutorial
Homology Modeling of Protein-Ligand Binding Sites with IFD-MD
Create a homology model of TYK2 from JAK3 and including a bound ligand. Compare this model with the crystal structure for TYK2 bound to 4GIH.
Training Videos
 Video
Life Science
Video
Life Science
Getting Going with Maestro BioLuminate
A free video series introducing the basics of using Maestro Bioluminate.
 Video
Life Science
Video
Life Science
- Video
Introducing Ligand Designer
An overview of the LigandDesigner workflow, Editing in 2D and 3D, using display options and overlays, and accessing the Admin Panel.
Publications
- Publication
- Dec 3, 2025
Glide WS: Methodology and Initial Assessment of Performance for Docking Accuracy and Virtual Screening
Friesner, et al. Journal of Chemical Theory and Computation, 2025, 21(24), 12696–12708
- Publication
- Oct 13, 2025
Accelerated in silico discovery of SGR-1505: A potent MALT1 allosteric inhibitor for the treatment of mature B-cell malignancies
Nie Z, et al. J. Med. Chem., 2025
- Publication
- Oct 12, 2025
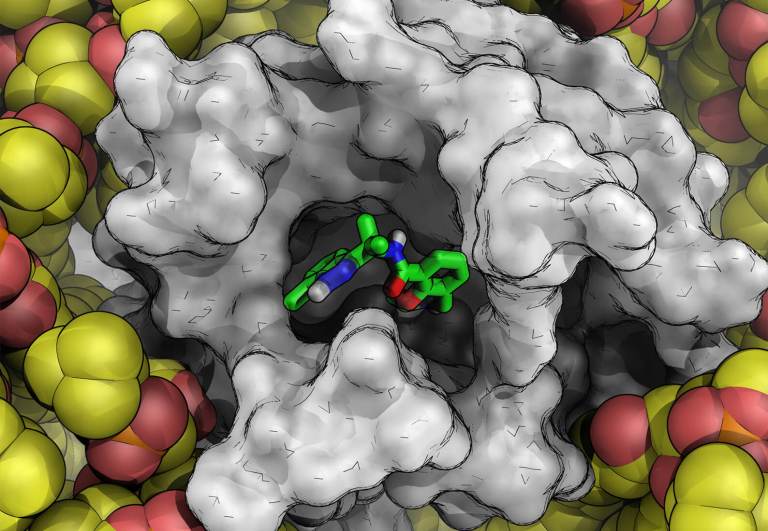
Discovery of highly potent noncovalent inhibitors of SARS-CoV-2 main protease through computer-aided drug design
Okabe A, et al. J Med Chem, 2025
Case Studies
 Case Study
Life Science
Materials Science
Case Study
Life Science
Materials Science
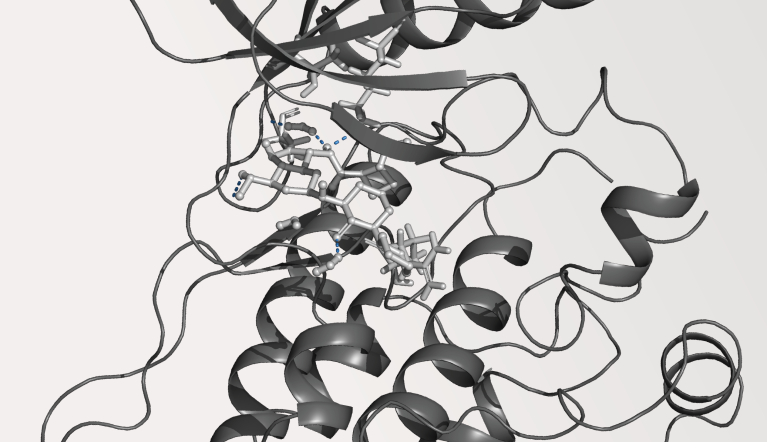 Case Study
Life Science
Case Study
Life Science
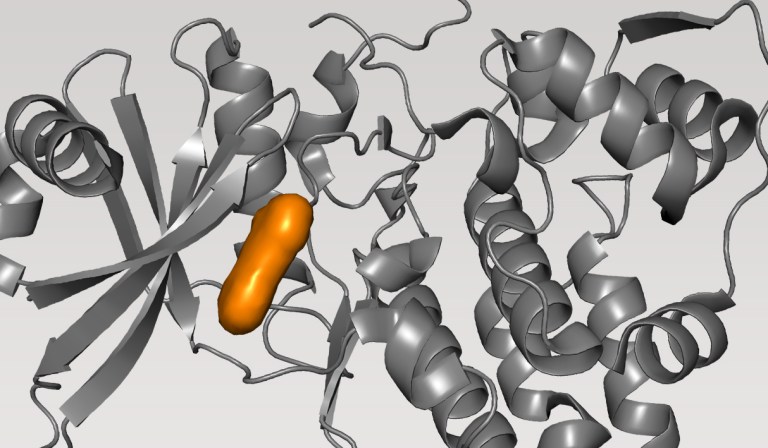 Case Study
Life Science
Case Study
Life Science
White Papers
 White Paper
Life Science
White Paper
Life Science
- Jan 29, 2026
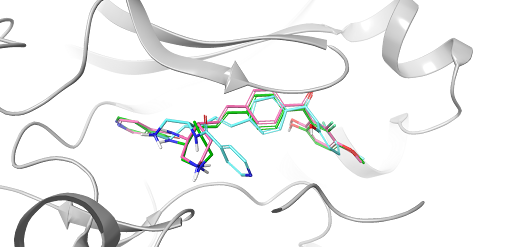
FEP+ Pose Builder — maximizing utility and productivity in FEP simulations
FEP+ Pose Builder is a methodological advancement introduced as an integrated feature to drastically enhance accessibility, user-friendliness, and productivity within the FEP+ pipeline.
 White Paper
Life Science
White Paper
Life Science
- Oct 29, 2024
20 Years of Glide: A Legacy of Docking Innovation and the Next Frontier with Glide WS
Glide has long set the gold standard for commercial molecular docking software due to its robust performance in both binding mode prediction and empirical scoring tasks, ease of use, and tight integration with Schrödinger’s Maestro interface and molecular discovery workflows.
 White Paper
Life Science
White Paper
Life Science
Quick Reference Sheets
- Quick Reference Sheet
Force Field Builder
A one-page guide to calculate missing torsion parameters for ligands using the Force Field Builder panel.
- Quick Reference Sheet
Ligand Interaction Diagram
A one-page guide to using the Ligand Interaction Diagram for examining ligand-receptor interactions.
- Quick Reference Sheet
GlideMap
A one-page guide to using the GlideMap GUI for ligand placement guided by experimental density.

Latest insights from Extrapolations blog
Training & Resources
Online certification courses
Level up your skill set with hands-on, online molecular modeling courses. These self-paced courses cover a range of scientific topics and include access to Schrödinger software and support.
Free learning resources
Learn how to deploy the technology and best practices of Schrödinger software for your project success. Find training resources, tutorials, quick start guides, videos, and more.