JUN 22, 2022
DFT calculations in materials science courses
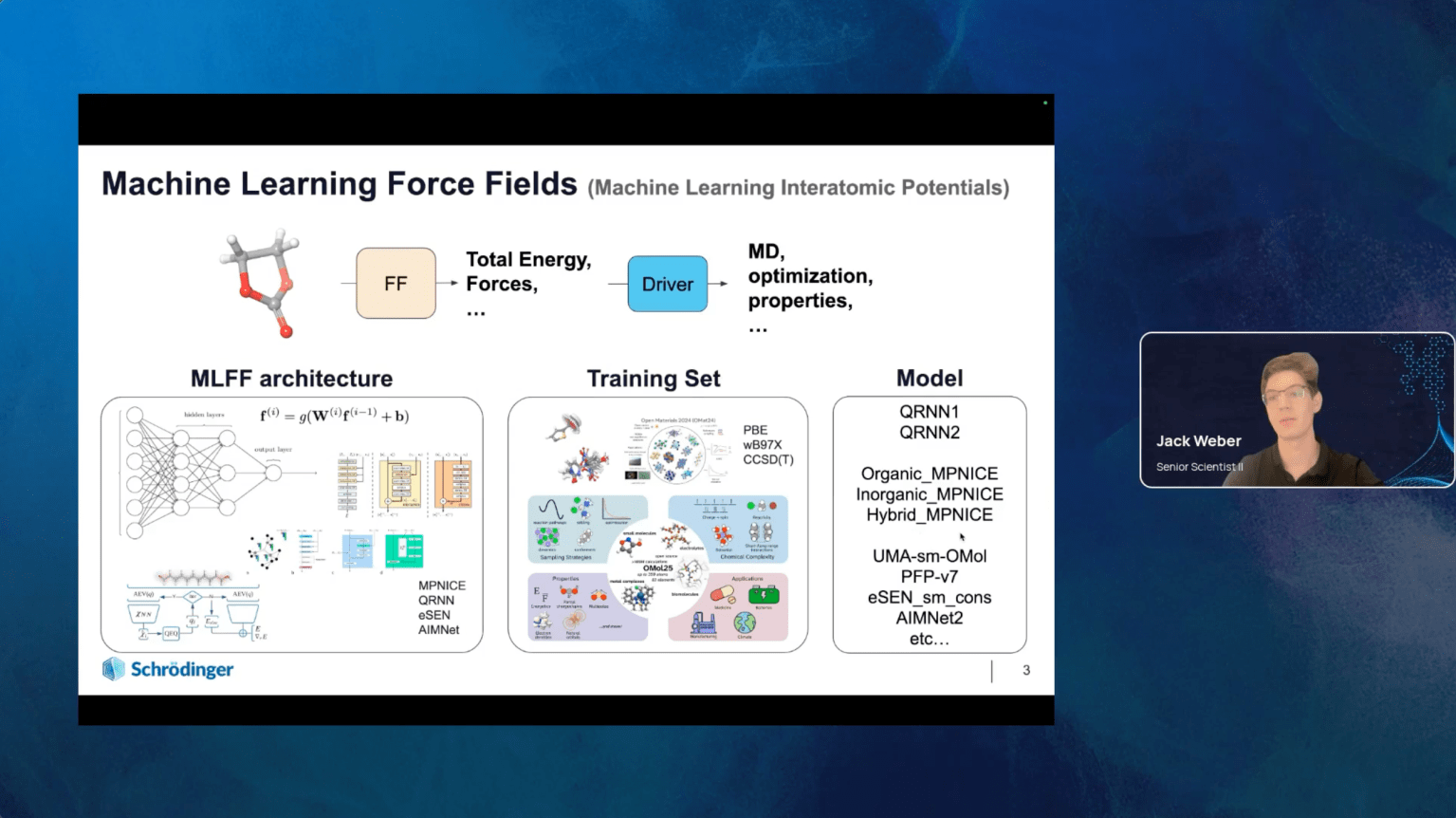
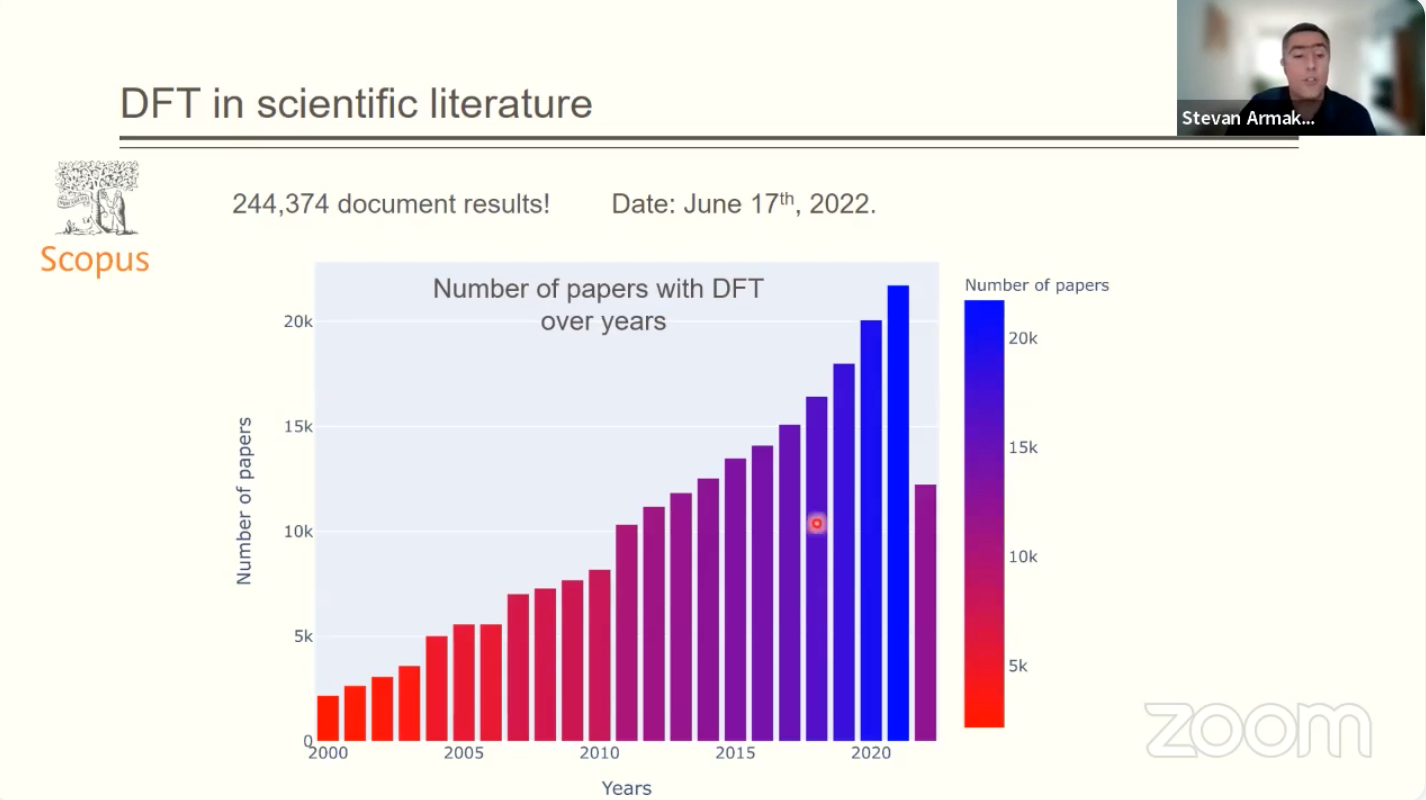
The development of computational methods for studying molecules and periodic structures has strongly influenced materials science. Over the years, computational studies have become the first step in developing novel materials. Many computational methods have become available, but the density functional theory (DFT) stood out due to the best cost-efficiency ratio.
Understanding the properties of both molecules and periodic structures is equally important in materials science, and the corresponding DFT methods have been developed. For convenience, it is imperative for users to perform both molecular and periodic DFT calculations in a single modeling package. Since the incorporation of the Quantum Espresso program, Schrödinger Materials Science Suite (SMSS) has become a unique modeling package covering both molecular and periodic DFT calculations. This move expanded the possibilities for Schrödinger users to investigate the periodic structures and allowed them to use SMSS to teach the fundamentals of computational condensed matter physics and address any relevant topic in materials science.
In this lecture it will be demonstrated how molecular and periodic DFT calculations available in SMSS can be used to teach selected concepts in materials science. The simple and intuitive graphical user interface of SMSS allows students to quickly adopt the fundamentals of materials modeling and the opportunity to engage in research activities with their professors. The experience from the recent training using the Teaching with Schrödinger platform at the University of Novi Sad will also be shared.
Our Speaker

Stevan Armakovic
University of Novi Sad
Stevan is a physicist with expertise in computational materials and molecular modeling. He applies atomistic calculations to understand structural, reactive, transport and optoelectronic properties of various molecules, (in)organic (nano)materials, ionic liquids, polymers, etc. He has a demonstrated history of applying a vast number of computational tools for atomistic calculations. He is currently affiliated with the Department of Physics, Faculty of Sciences of the University of Novi Sad, where he teaches various physics courses. Stevan is also a high-school professor, teaching “Atomic and molecular physics” and “Modeling in physics” to students with exceptional abilities in physics at the “Jovan Jovanović Zmaj” high school. Stevan has published more than 120 papers in journals with impact factors, and he regularly involves high school and faculty students in his research activities. He is a verified peer reviewer at Publons, with more than 300 pre-publication reviews for journals with impact factors. He serves as a Board President of an NGO dedicated to the international development of academic and scientific collaboration (www.aidasco.org).