MAY 29, 2024
Virtual Biotechs – A Student Group Learning Project Enhanced Through Application of Schrödinger Maestro in a Medicinal Chemistry Course
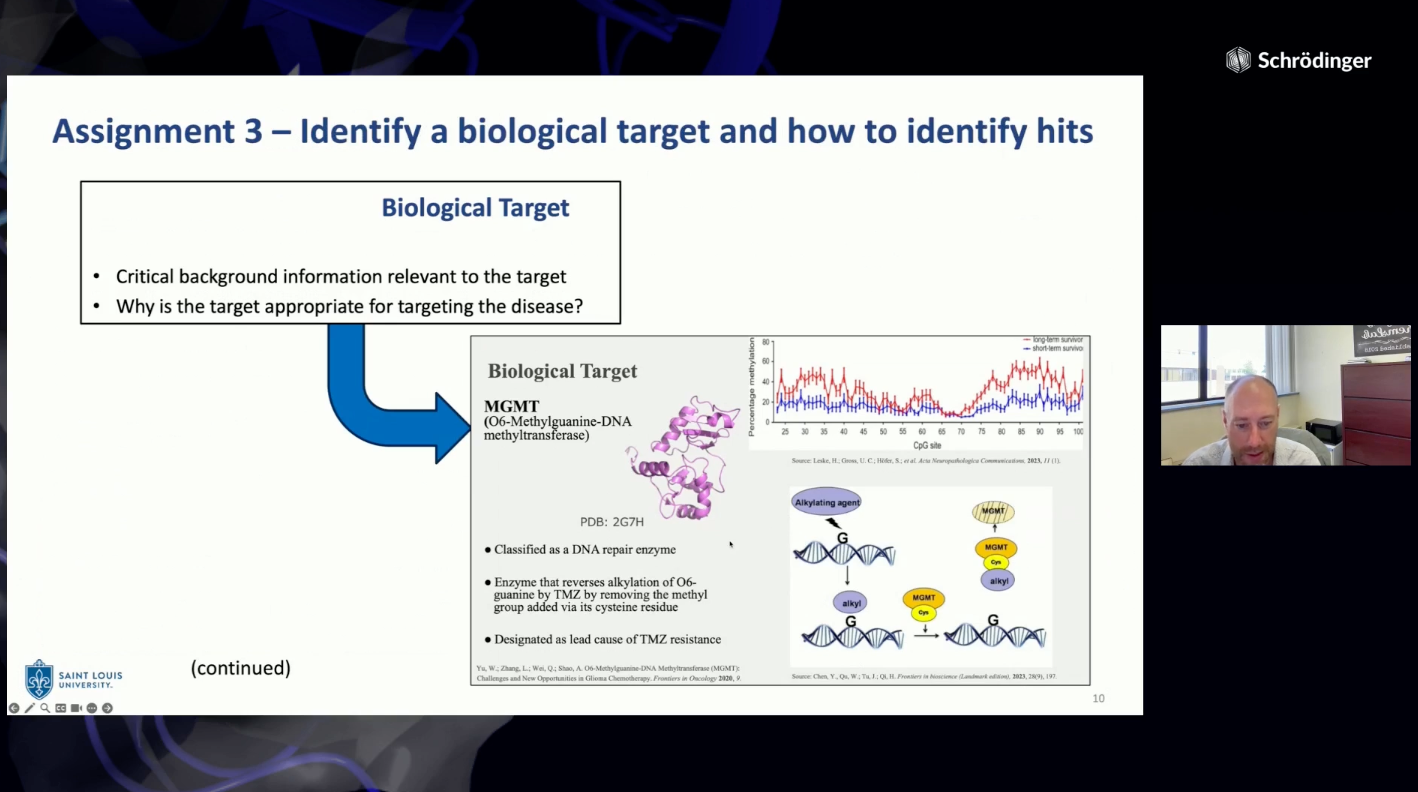
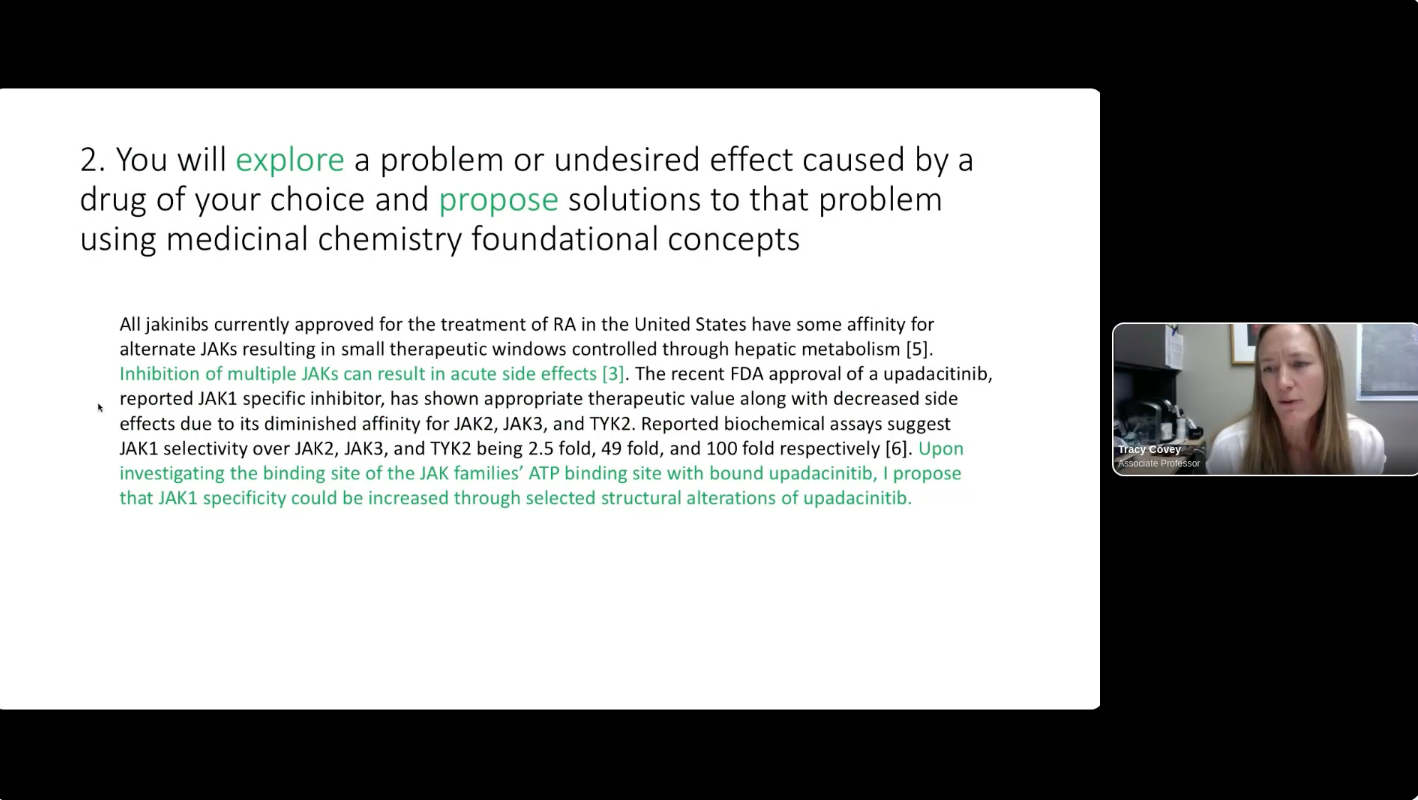
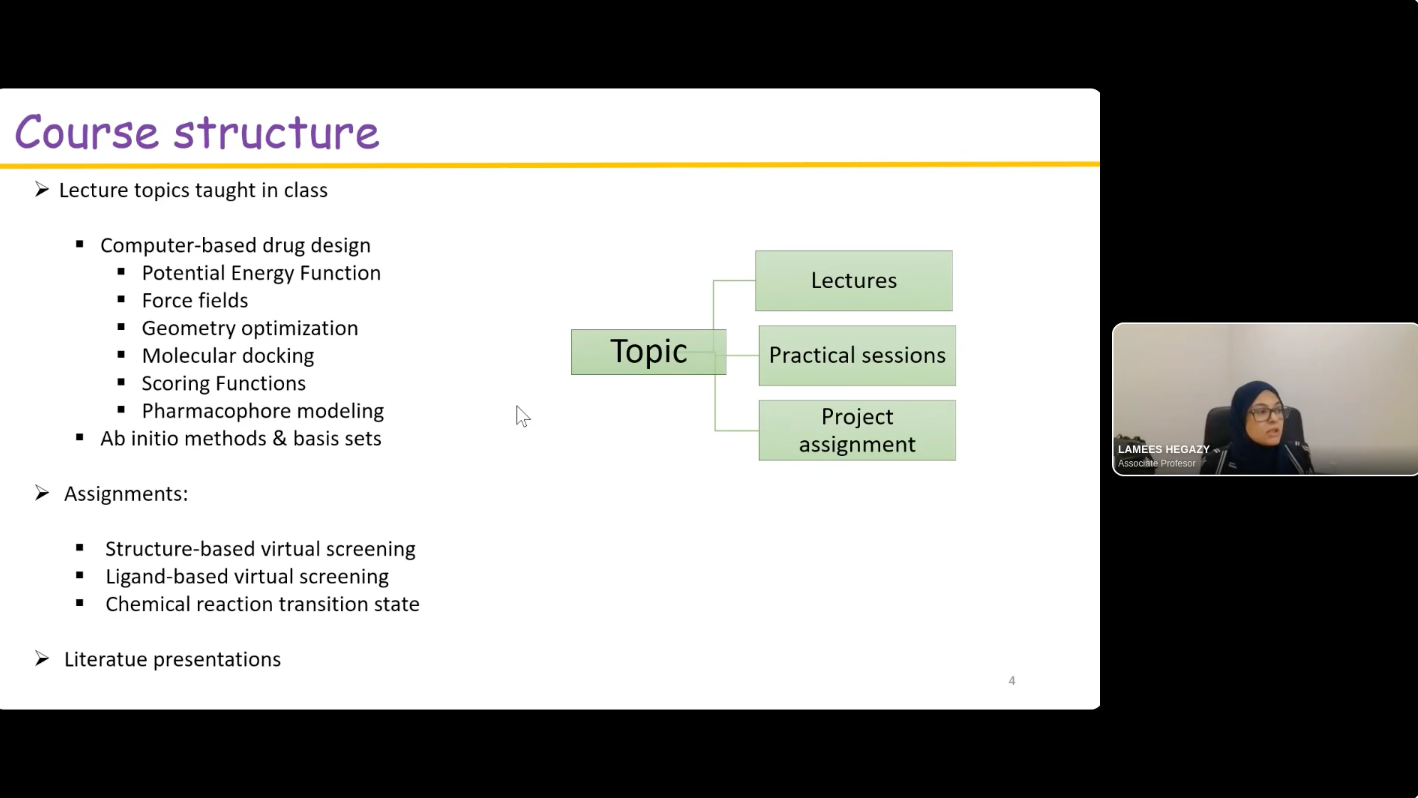
The Department of Chemistry at Saint Louis University offers an introductory medicinal chemistry course for juniors, seniors, and graduate students every spring. In 2019, an active learning group project was developed over the course of the semester wherein student teams of “virtual biotech” companies were formed to work through the drug discovery process on a virtual project. This was introduced as an experiment to help break up a 2.5 hour night course while applying what the students were learning in a more “real world” application that mimics working on diverse teams that occurs naturally in the pharmaceutical industry workforce. Student groups work to identify a disease with unmet medical need, target candidate profile, suitable biological targets for drug discovery, processes for hit identification, and propose an optimization strategy. The group project continues to evolve each year based on student feedback. Incorporation of structure-based drug design utilizing Schrödinger Maestro molecular modeling software was introduced in 2022 and is now integrated throughout the semester in individual weekly assignments to learn the software and the group project to utilize the software to apply what the students were learning in class to their project. The project teams conduct a virtual screen for their target using commercial screening libraries, evaluate hits, and use Ligand Designer to propose new analogs. At the end of the semester, the project teams present their work as a 10-minute project pitch to “virtual investors” (their classmates) to select a project for “virtual investment”.
Our Speaker

Marvin Meyers
Saint Louis University
Meyers is an experienced medicinal chemist and drug discovery scientist. Upon completion of his PhD in Chemistry at the University of Illinois-Urbana Champaign, he spent a decade at Pharmacia and Pfizer working on new drug discovery for a variety of diseases resulting in two novel compounds entering human clinical trials. Since joining Saint Louis University in 2010, his research focus shifted to the design and synthesis of novel drug candidates for rare and infectious diseases, focusing on parasitic, fungal, bacterial and viral diseases with few, if any, treatment options. This work is highly collaborative where the Meyers lab uses organic synthesis to develop structure-activity relationships on lead molecules in partnership with leading disease experts towards the goal of identifying drug candidates for clinical trials. These efforts are supported by grants from the National Institutes of Health and the SLU Research Institute. His work has resulted in 61 peer-reviewed publications, 34 patent applications and 7 issued US patents. In 2021, he was elected as a Senior Member of the National Academy of Inventors.