AUG 6, 2025
Advancing machine learning force fields for materials science applications
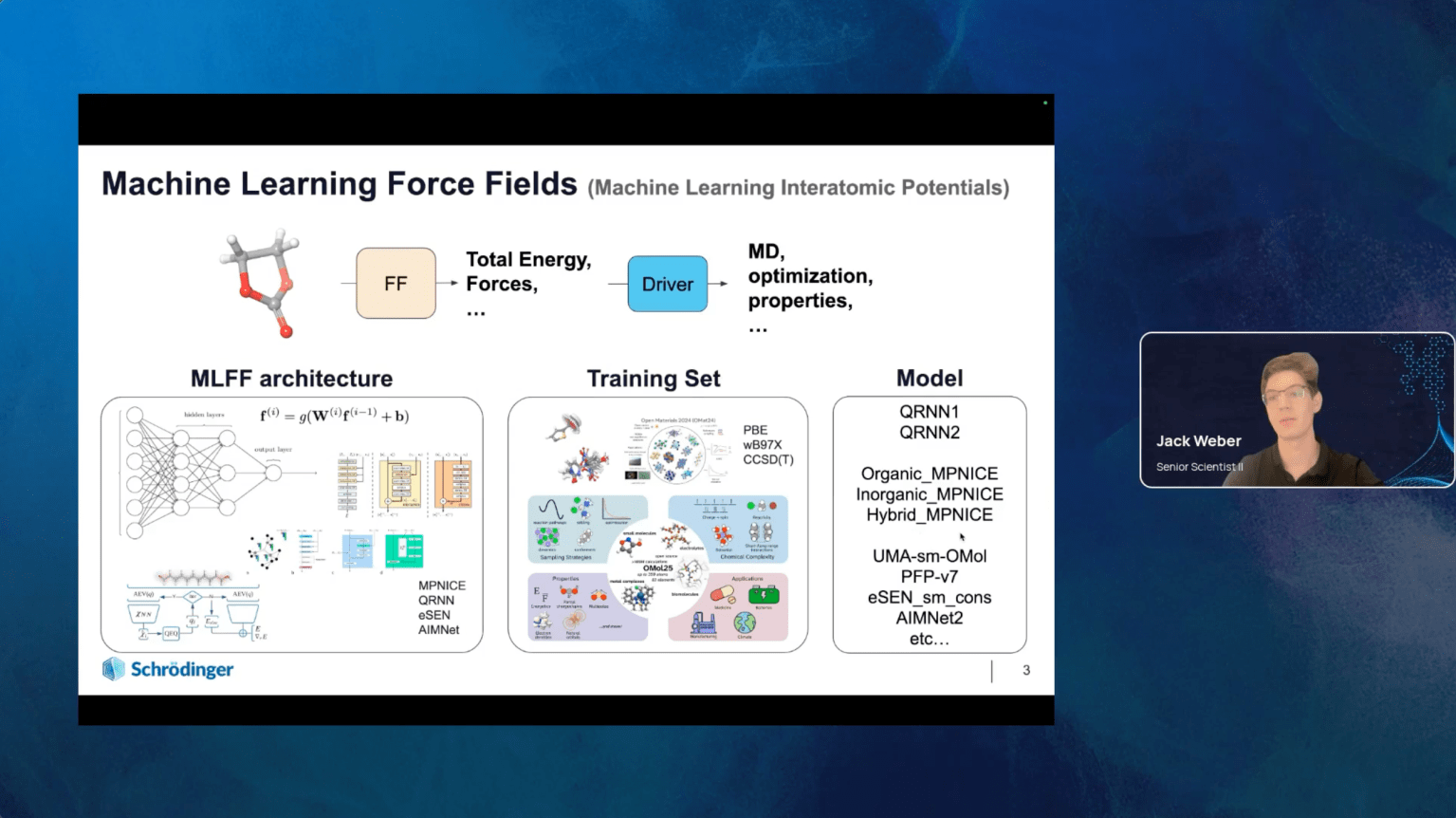
Machine learning force fields (MLFFs), also referred to as machine learning interatomic potentials, have emerged as a critical tool for the cost-efficient atomistic simulations of diverse chemical systems, often achieving density functional theory (DFT) accuracy at a fraction of the cost. Recent advances in message passing networks have removed the drawback of previous MLFFs that were limited by the number of unique atomic elements they could model. Furthermore, inclusion of atomic charges and electrostatics through charge equilibration approaches have enabled accurate representations of multiple charge states, ionic systems, and electronic response properties, while simultaneously improving accuracy using explicit long-range interactions.
In this webinar, we will introduce Schrödinger’s state-of-the-art MLFF architecture, called Message Passing Network with Iterative Charge Equilibration (MPNICE), which incorporates explicit electrostatics for accurate charge representations. We present a family of pre-trained models trained on materials covering the entire periodic table (89 elements). MPNICE prioritizes efficient throughput, enabling atomistic simulations at length and time scales that were previously inaccessible without sacrificing accuracy. We will outline available tools in the Schrödinger suite that incorporate MLFFs to enable larger and more complex simulations for materials design, providing industry relevant examples throughout.
Highlights:
- Overview of MPNICE – a message passing MLFF architecture that includes atomic partial charges and long-range interactions, while maintaining speeds an order of magnitude faster than comparable models
- Highlights of recent applications of MPNICE, including general organic, inorganic, and hybrid (organic and inorganic) models to address industry relevant needs
Our Speaker

Jack Weber
Senior Scientist, Schrödinger
Jack Weber is a Senior Scientist at Schrödinger, where he develops machine learning force fields (MLFFs) for applications in drug discovery and materials science. Jack received his PhD in Chemical Physics in 2023 from Columbia University, advised by Professors Richard Friesner and David Reichman. In his doctoral research, he used advanced computational methods to investigate fundamental problems in chemistry and materials science, including improving ab-initio methods in electronic structure to treat strongly correlated systems.















What our alumni say