- Publication
- Feb 2, 2011
Further Characterization of the [Fe-Fe]-Hydrogenase Maturase’ HydG
Tron, et al. Eur. J. Inorg. Chem., 2011, 7, 1121-1127
- Publication
- Jan 19, 2011
Carbon Monoxide Dehydrogenase Reaction Mechanism: A Likely Case of Abnormal CO2 Insertion to a Ni-H- Bond
Amara, et al. Inorg. Chem., 2011, 50, 1868-1878
- Publication
- Jan 11, 2011
Combined receptor and ligand-based approach to the universal pharmacophore model development for studies of drug blockade to the hERG1 pore domain
Durdagi, et al. J. Chem. Inf. Model., 2011, 51, 463-74
- Publication
- Jan 4, 2011
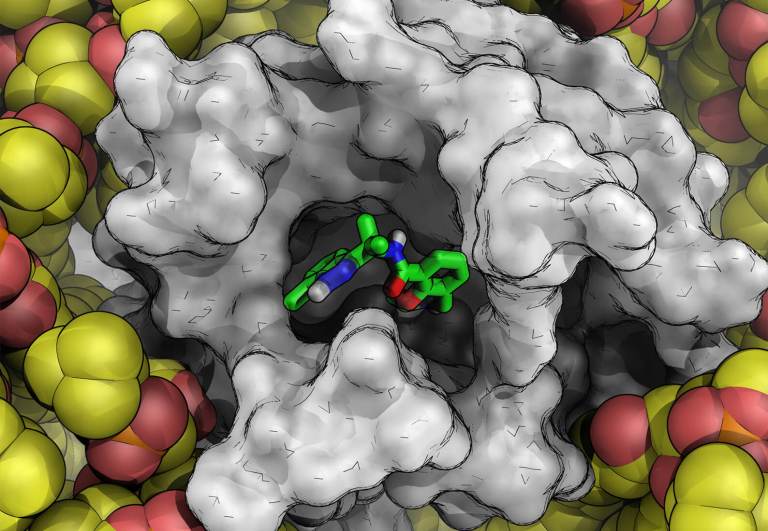
Ligand binding to protein-binding pockets with wet and dry regions
Wang, et al. Proc. Natl. Acad. Sci. U S A., 2011, 108, 1326-30
- Publication
- Jan 1, 2011
Computational Insights into Binding of Bisphosphates to Farnesyl Pyrophosphate Synthase
Ohno, et al. Curr. Med. Chem., 2011, 18, 220-233
- Publication
- Dec 21, 2010
Structural basis for effectiveness of siderophore-conjugated monocarbams against clinically relevant strains of Pseudomonas aeruginosa.
Han, et al. Proc. Natl. Acad. Sci. U S A., 2010, 107, 22002-7
- Publication
- Oct 18, 2010
Unexpected Cleavage of 2-Azido-2-(hydroxymethyl)oxetanes: Conformation Determines Reaction Pathway-
Farber, et al. J. Org. Chem., 2010, 75, 7565-7572
- Publication
- Oct 15, 2010
Conformationally Restricted Homotryptamines. Part 7: 3-cis-(3-Aminocyclopentyl)indoles As Potent Selective Serotonin Reuptake Inhibitors
King, et al. J. Med. Chem., 2010, 53, 7564-7572
- Publication
- Sep 24, 2010
NADH Oxidase Activity of Bacillus subtilis Nitroreductase NfrA1: Insight into its Biological Role
Cortial, et al. FEBS Lett., 2010, 584, 3916-22
- Publication
- Sep 22, 2010
Estimating binding affinities by docking/scoring methods using variable protonation states
Park, et al. Proteins, 2011, 79, 304-314
- Publication
- Sep 1, 2010
Analysis and Comparison of 2D Fingerprints: Insights into Database Screening Performance Using Eight Fingerprint Methods
Duan, et al. J. Molec. Graph. Model., 2010, 29, 157-170
- Publication
- Sep 1, 2010
New hypotheses about the structure-function of proprotein convertase subtilisin/kexin type 9: Analysis of the epidermal growth factor-like repeat A docking site using WaterMap
Pearlstein, et al. Proteins, 2010, 78, 2571-2586
Events
 Event
Life Science
Event
Life Science
- Mar 10, 2026
38th Molecular Modeling Workshop 2026
Schrödinger is excited to be participating in the 38th Molecular Modeling Workshop 2026 conference taking place on March 9th – 10th in Erlangen, Germany.
 Event
Life Science
Event
Life Science
- Mar 10, 2026
AI in drug discovery 2026
Schrödinger is excited to be participating in the AI in drug discovery 2026 conference taking place on March 9th – 10th in London, United Kingdom.
 Webinar
Materials Science
Webinar
Materials Science
- Mar 10, 2026
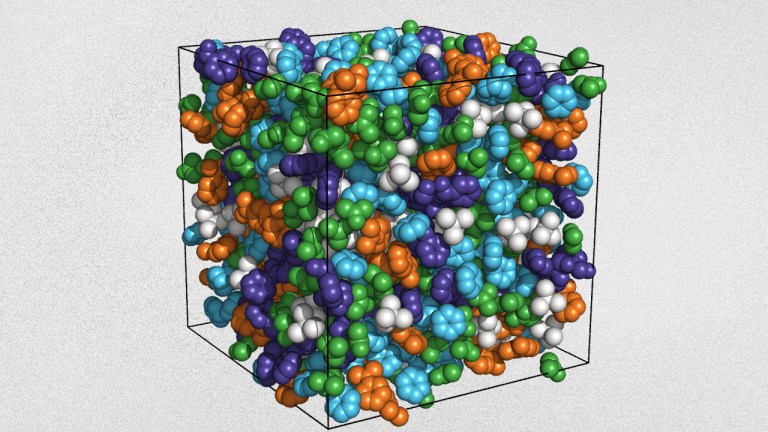
Formulation ML and Optimization: Making advanced property prediction and experimental design fast and accessible
We will showcase how easy it is to apply these tools using experimental datasets across broad MS applications, including formulations, consumer goods, batteries, pharmaceuticals, and beyond.
Webinars
 Webinar
Life Science
Webinar
Life Science
- Mar 18, 2026
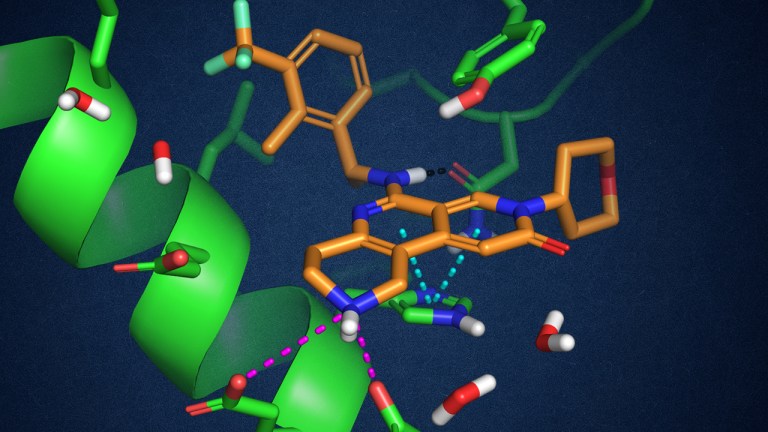
デジタル創薬セミナー ~計算化学がもたらす創薬プロセスの変貌~ 第23回
Rethinking the rules: Exploiting solvent exposed salt-bridge interactions with free energy perturbation simulations for the discovery of potent inhibitors of SOS1
 Webinar
Life Science
Webinar
Life Science
- Mar 19, 2026
Introducing RetroSynth: Breaking the synthesis bottleneck with AI and physics-based modeling
Join us for the introduction of RetroSynth, Schrödinger’s AI-driven synthesis planning platform. RetroSynth is engineered to accelerate and scale conventional retrosynthesis by harnessing advanced deep learning algorithms.
 Webinar
Life Science
Webinar
Life Science
- Mar 26, 2026
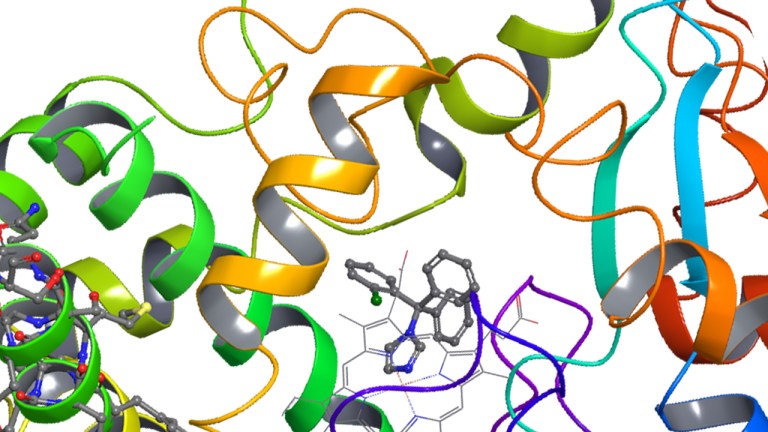
Embracing a new era of toxicity screening: Atomic-resolution modeling to mitigate off-target liabilities
Join us for a technical overview of Schrödinger’s Predictive Toxicology solution. This session will demonstrate how physics-based, atomic-resolution modeling transforms toxicology from a reactive “filter” into a proactive “design tool.”
Documentation
- Documentation
Learning Path: Oligonucleotide Modeling
A structured overview of tools and workflows for nucleic acids in drug discovery.
- Documentation
WaterMap
Efficiently converged MD simulations are run with explicit water molecules, and resultant trajectories are analyzed to cluster hydration sites.
- Documentation
SiteMap
Identify binding sites, including allosteric binding sites and protein-protein interfaces, and evaluate their druggability.
Tutorials
- Tutorial
Structure-Based Virtual Screening using Glide
Prepare receptor grids for docking, dock molecules and examine the docked poses.
- Tutorial
Ligand Binding Pose Generation for FEP+
Generate starting poses for FEP simulations for a series of BACE1 inhibitors using core constrained docking.
- Tutorial
Homology Modeling of Protein-Ligand Binding Sites with IFD-MD
Create a homology model of TYK2 from JAK3 and including a bound ligand. Compare this model with the crystal structure for TYK2 bound to 4GIH.
Training Videos
 Video
Life Science
Video
Life Science
Getting Going with Maestro BioLuminate
A free video series introducing the basics of using Maestro Bioluminate.
 Video
Life Science
Video
Life Science
- Video
Introducing Ligand Designer
An overview of the LigandDesigner workflow, Editing in 2D and 3D, using display options and overlays, and accessing the Admin Panel.
Publications
- Publication
- Oct 13, 2025
Accelerated in silico discovery of SGR-1505: A potent MALT1 allosteric inhibitor for the treatment of mature B-cell malignancies
Nie Z, et al. J. Med. Chem., 2025
- Publication
- Oct 12, 2025
Discovery of highly potent noncovalent inhibitors of SARS-CoV-2 main protease through computer-aided drug design
Okabe A, et al. J Med Chem, 2025
- Publication
- Sep 17, 2025
Accurate hydration free energy calculations for diverse organic molecules with a machine learning force field
Xie, et al. ChemRxiv, 2025, 1, Preprint
Case Studies
 Case Study
Life Science
Materials Science
Case Study
Life Science
Materials Science
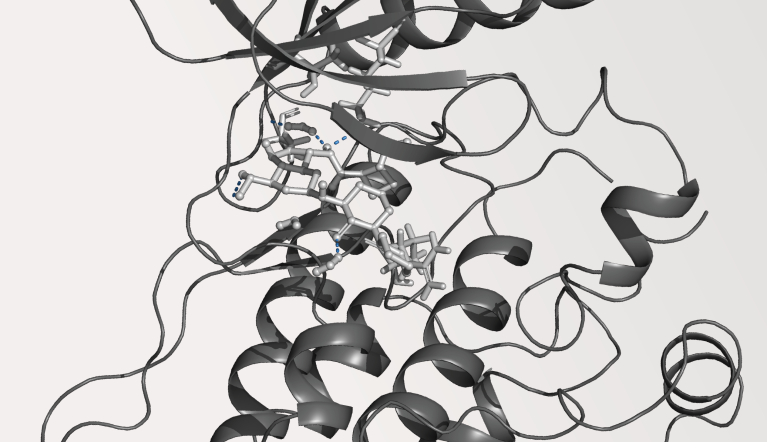 Case Study
Life Science
Case Study
Life Science
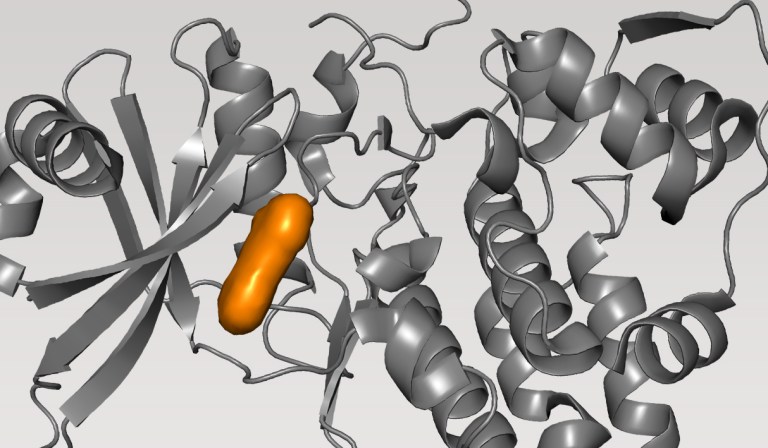 Case Study
Life Science
Case Study
Life Science
White Papers
 White Paper
Life Science
White Paper
Life Science
- Jan 29, 2026
FEP+ Pose Builder — maximizing utility and productivity in FEP simulations
FEP+ Pose Builder is a methodological advancement introduced as an integrated feature to drastically enhance accessibility, user-friendliness, and productivity within the FEP+ pipeline.
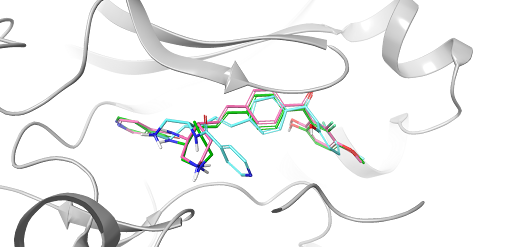 White Paper
Life Science
White Paper
Life Science
- Oct 29, 2024
20 Years of Glide: A Legacy of Docking Innovation and the Next Frontier with Glide WS
Glide has long set the gold standard for commercial molecular docking software due to its robust performance in both binding mode prediction and empirical scoring tasks, ease of use, and tight integration with Schrödinger’s Maestro interface and molecular discovery workflows.
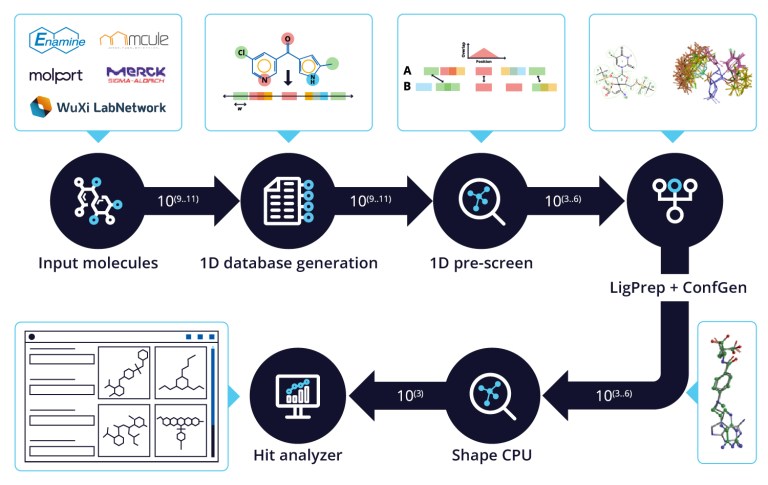 White Paper
Life Science
White Paper
Life Science
Quick Reference Sheets
- Quick Reference Sheet
Force Field Builder
A one-page guide to calculate missing torsion parameters for ligands using the Force Field Builder panel.
- Quick Reference Sheet
Ligand Interaction Diagram
A one-page guide to using the Ligand Interaction Diagram for examining ligand-receptor interactions.
- Quick Reference Sheet
GlideMap
A one-page guide to using the GlideMap GUI for ligand placement guided by experimental density.

Latest insights from Extrapolations blog
Training & Resources
Online certification courses
Level up your skill set with hands-on, online molecular modeling courses. These self-paced courses cover a range of scientific topics and include access to Schrödinger software and support.
Free learning resources
Learn how to deploy the technology and best practices of Schrödinger software for your project success. Find training resources, tutorials, quick start guides, videos, and more.