MAR 18, 2026
The Importance of Human Know-How in AI Execution for Materials R&D
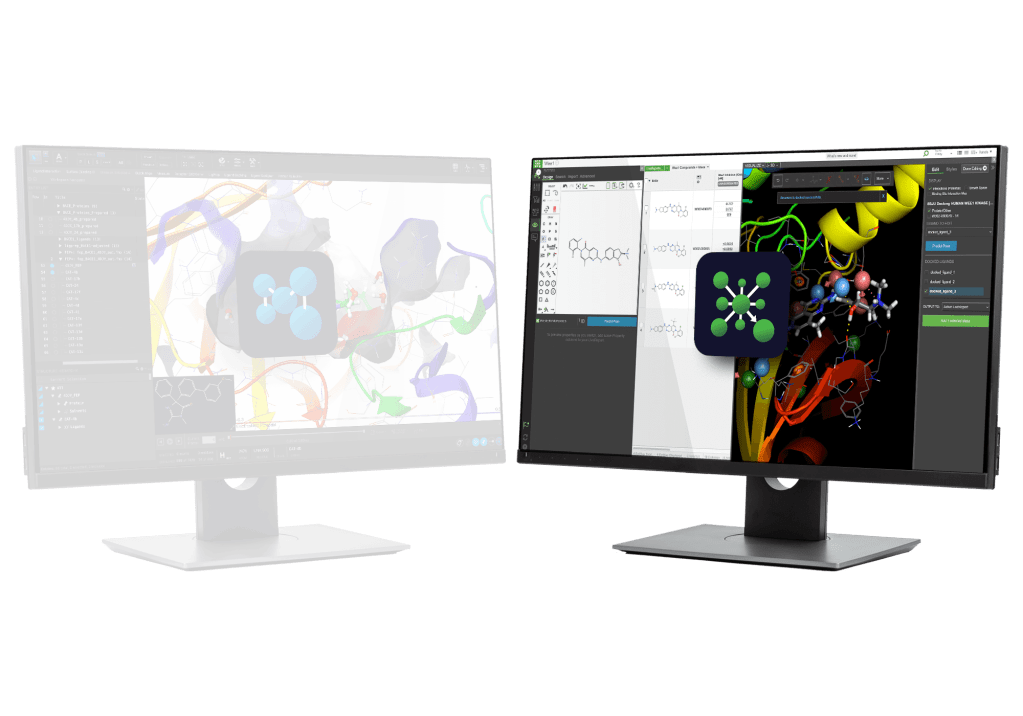
AI and machine learning (ML) are often sold as push-button solutions for materials design and discovery, but they lack value without a rigorous foundation. While the current AI revolution provides unprecedented speed and possibilities, human know-how is a key ingredient for ensuring complex methods lead to high impact outcomes. True innovation happens at the intersection of physics-based simulation, AI/ML, and human expertise. Join us to explore how Schrödinger’s domain experts integrate these three pillars to streamline material optimization.
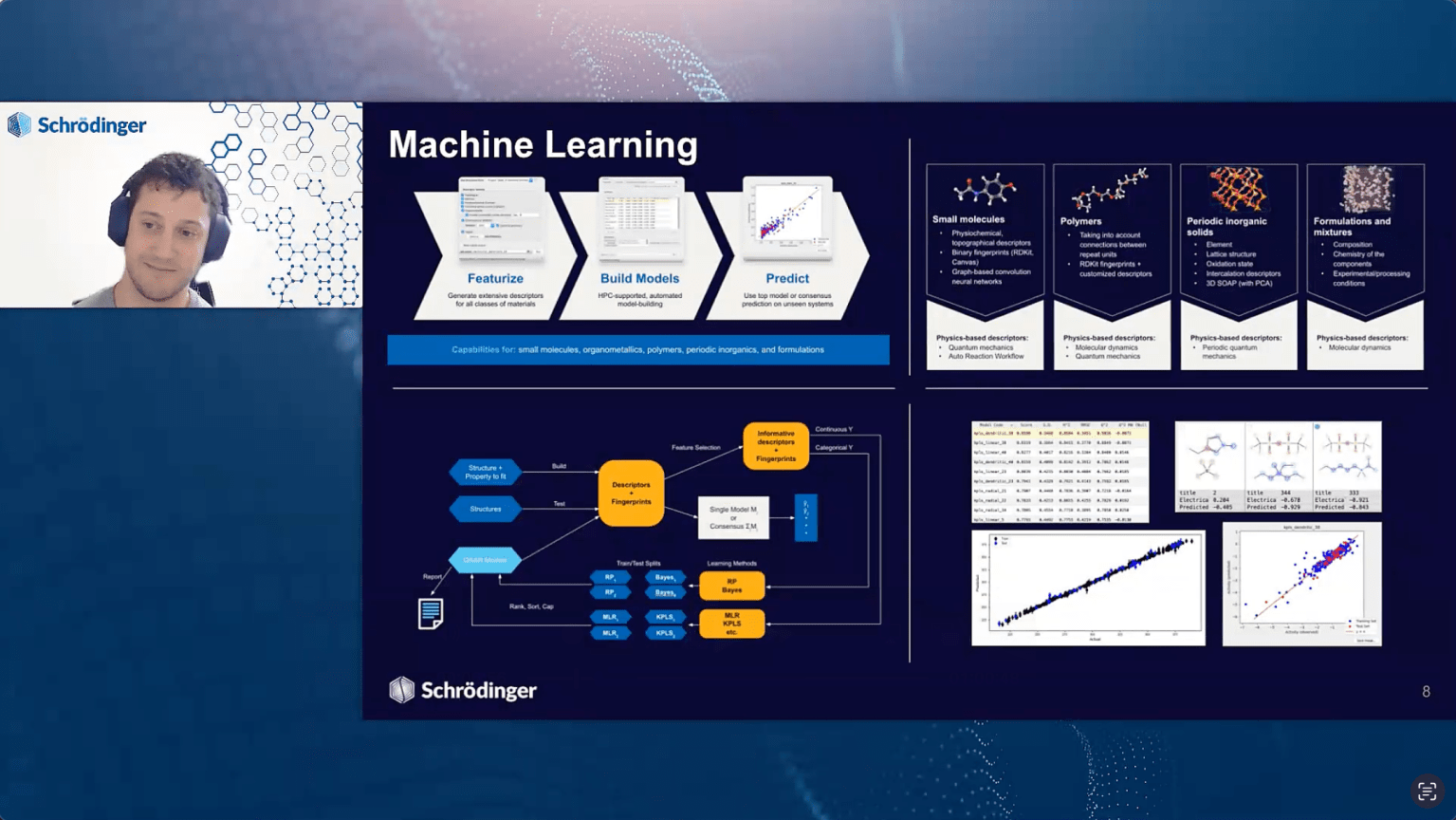
We’ll discuss how to move beyond the hype and apply digital chemistry strategies that deliver meaningful business results. We will introduce Schrödinger’s Materials Science platform, and share high-impact case studies from a variety of industries and applications, ranging from small molecules to formulations and electronics to industrials. Recent advancements, such as device level ML and cutting-edge machine learning force field (MLFF) architectures will be presented.
Key Learning Objectives:
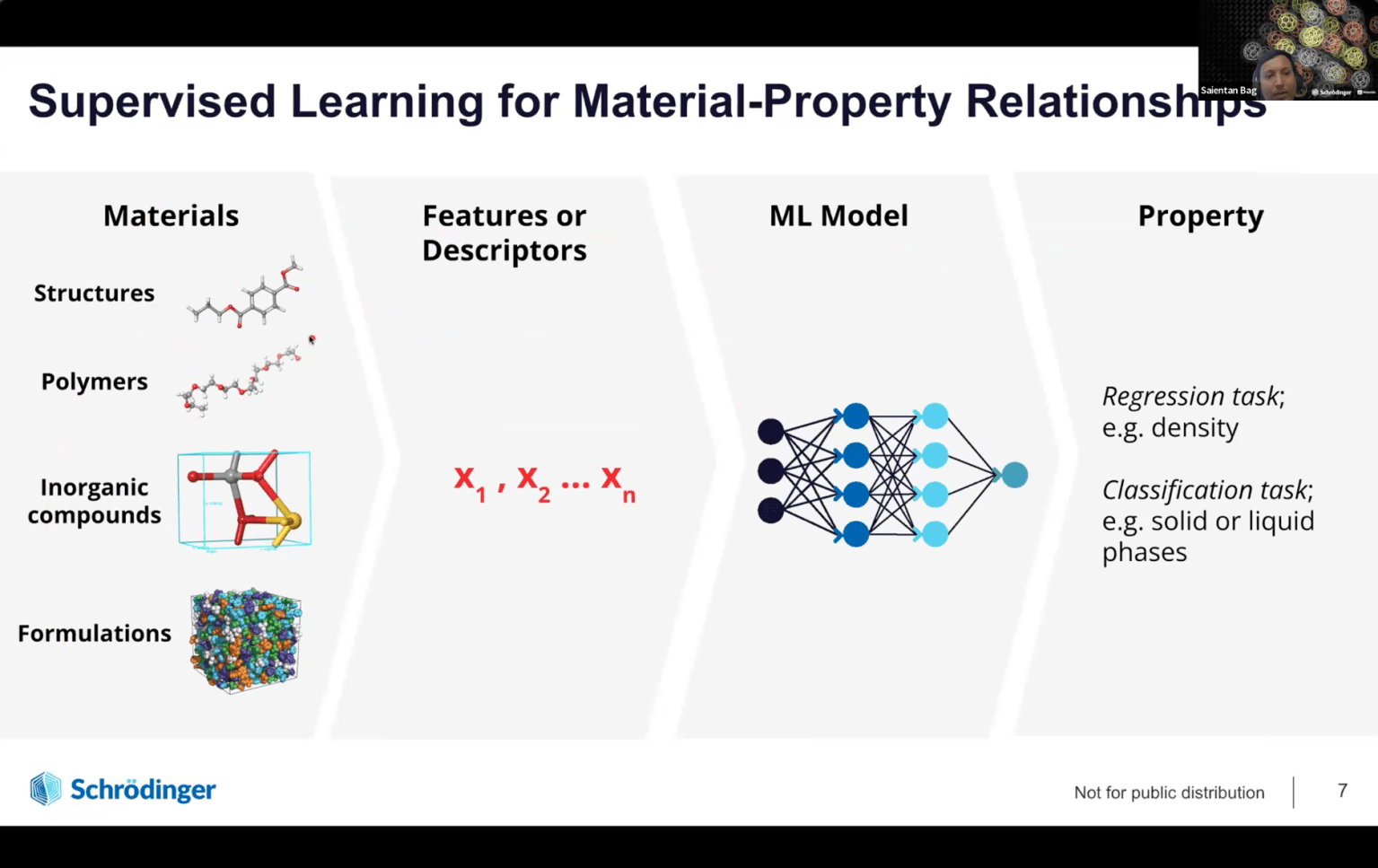
- Why physics-based modeling is essential to complement AI/ML predictions
- Real-world applications where digital chemistry has reduced discovery timelines from years to months across industries
- How our expert-led support ensures project success for modeling novices and veterans alike
Who Should Attend:
- R&D Leaders
- Innovation Managers
- Digitization Managers
- Synthetic Chemists
- Materials Scientists
- Chemical Engineers
- Materials Research Engineers
- Computational Chemists
- Computational Materials Scientists
Our Speaker

Michael Rauch
Director of Materials Science, Schrödinger
Michael Rauch is a Director at Schrödinger specializing in materials science and education. Michael earned his Ph.D. from Columbia University in synthetic organometallic chemistry as an NSF Graduate Research Fellow before pursuing a postdoctoral role in organic chemistry at the Weizmann Institute of Science as a Zuckerman Postdoctoral Scholar. Michael is particularly interested in green, sustainable chemistry and transforming the way that synthetic chemists utilize molecular modeling via practical education.