- October 13th, 2025
- 10:30 – 15:30 BST
- Cambridge, United Kingdom


Informatics for Medicinal Chemists
Dear Medicinal Chemists,
Ever feel your DMTA cycles are not advancing as quickly as you’d like? Is it a challenge to bring all your data from in silico predictions to experimental results into one central view to quickly decide what to design next? Effectively sharing hypotheses with your team and securely tracking data with CRO partners present their own sets of challenges.
Schrödinger therefore invites you to a specialized and free-of-charge “Lunch & Learn” workshop on Monday, October 13th at the Clayton Hotel Cambridge, designed to tackle these exact workflows and collaboration challenges.
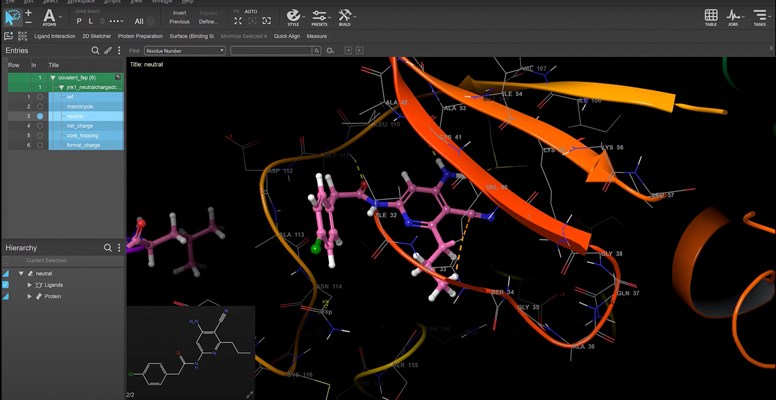
We’ll be diving deep into our informatics platform, LiveDesign, to show you how all members of a drug design team can work together to solve these challenges. At this event, Schrödinger will host a hands-on CDK2 inhibitor design challenge where you’ll be able to use LiveDesign.
Date & Time:
Monday, October 13, 2025
From 10:30 to 15:30 BST
Program:
Part 1: Welcome Coffee and Introductory Talk about the Platform and Success Stories
10:30 – 12:00 BST
Olivia Lynes, Senior Strategic Deployment Manager II, Enterprise Informatics
+ Lunch
12:00 – 13:00 BST
Part 2: Workshop and Design Challenge on CDK2 Inhibitor with LiveDesign
13:00 – 14:30 BST
Olivia Lynes, Senior Strategic Deployment Manager II, Enterprise Informatics
Hands-on CDK2 inhibitor design challenge where you’ll be able to use LiveDesign on:
- Predicted physicochemical properties.
- Machine learning models.
- Ligand Designer: a validated docking and design model.
- A tool which searches ChEMBL and vendor databases (with over 1 billion total compounds) at rapid speeds to estimate the novelty of designed compounds.
- A Target Product Profile MPO.
Part 3: Interactive Q&A and Networking Session
14:30 – 15:30 BST
The afternoon session will feature a Q&A and networking session, providing an opportunity to present your questions and challenges, which the Schrödinger team will endeavor to address.
You can either join for the whole event or solely for the presentation session. All you need to bring is a laptop – no software installation is required. During the workshop, lunch will be served. The afternoon will feature an interactive Q&A and networking session, providing an opportunity to present your questions and challenges, which the Schrödinger team will endeavor to address.
We look forward to seeing you in Cambridge!
Register today to secure your seat!
The workshop is free to attend but preregistration is required as seats are limited. Previous-experience with the Schrödinger suite is not required.