SCC78 2024
- December 11th-13th, 2024
- Los Angeles, California
Schrödinger is excited to be participating in the SCC78 conference taking place on December 11th – 13th in Los Angeles, California. Join us for a presentation by Haidong Liu, Senior Scientist at Schrödinger, titled “Screening Antioxidant Ingredients Using Machine Learning and Physics-based Modeling .”
Screening Antioxidant Ingredients Using Machine Learning and Physics-based Modeling
Speaker:
Haidong Liu, Senior Scientist, Schrödinger
Abstract:
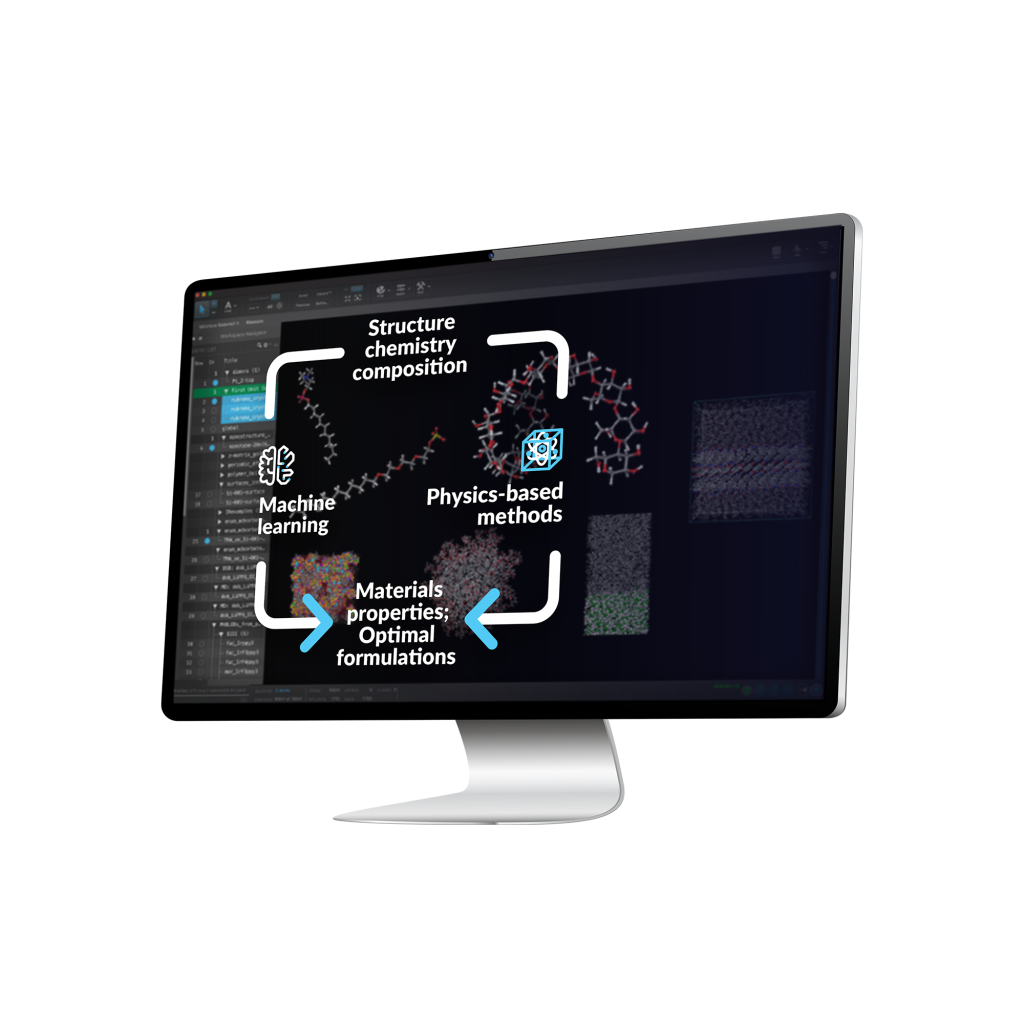
Antioxidants are an important ingredient for cosmetic products to alleviate oxidative stress. While high-throughput screening for new antioxidant candidates still remains challenging experimentally. And the data-driven machine learning models would require the input of a reliable dataset. Here we present an efficient computational approach that combines the physics-based and machine learning tools to address this issue, and this
approach only uses molecular structures as inputs.
We used molecular quantum mechanical (QM) calculation and machine learning to predict the antioxidant activity through hydrogen atom transfer (HAT) mechanism. We first constructed a library of flavonoid structures and then calculated the hydrogen dissociation energies of the hydroxyl group in solvents using QM. The machine learning model was trained and validated using the hydrogen dissociation energies from QM calculations. We can easily screen thousands of molecules, and this physics-based and machine learning combined approach can be used for other properties.

















Recent Testimonials