Schrödinger User Group Meeting – Materials Science Japan 2024
- October 3rd, 2024
- Tokyo, Japan
本会は、10月3日(木)、会場で開催いたします。
弊社サイエンティストや各製品の開発責任者から、最新機能、応用事例、今後の展望などを、セミナー形式でご紹介いたします。
さらに、パナソニックインダストリー株式会社 松澤 伸行様 による特別招待講演を予定しております。松澤様は、高度専門職 であるエグゼクティブエンジニアとして、各種電子部品用の材料設計・開発を担当されています。特別講演では、シュレーディンガーとのコラボレーションによる新規材料の開発事例をご紹介いただきます。
皆様のご参加を心よりお待ち申し上げております。
各発表のアブストラクトはこちらからご覧いただけます。
ご挨拶
分子動力学計算・密度汎関数計算と機械学習による電解液分子設計
パナソニックインダストリー株式会社 技術本部 プロセスデバイス革新センター エグゼクティブエンジニア 松澤 伸行様
Advancing Materials Science with Schrödinger: Latest Innovations, Future Roadmap, and Emerging Applications for Organic Electronics and Beyond
Mathew D. Halls, Senior Vice President, Materials Science, Schrödinger
Accelerating Cosmetics Design through Formulations Modeling at the Molecular Level
Jeffrey Sanders, Product Manager and Scientific Lead of Consumer Goods, Schrödinger
ランチ
Local Formal Chargeを用いた新規チタン酸窒化物の探索
シュレーディンガー株式会社 シニア サイエンティスト 青木 祐太
Combined Physics-Based and Machine Learning Approaches in the Design of Complex Materials
Anand Chandrasekaran, Product Manager Materials Science Informatics, Schrödinger
ニューラルネットワークポテンシャル表面上の強化学習による遷移状態
シュレーディンガー株式会社 シニア サイエンティスト 大塚 勇起
Automated Identification of Chemical Reaction Products with Nanoreactor
Pavel A. Dub, Senior Principal Scientist, Schrödinger
高速分子動力学計算と量子力学計算の組み合わせによるGHz以上の高周波数における誘電率と誘電正接の予測
シュレーディンガー株式会社 マテリアルズサイエンス シニア ディレクター 森里 嗣生
Polymers and Formulations in Schrödinger Materials Science Suite: From electronics to pharmaceuticals, new properties bring new applications
Andrea Browning, Director – Polymers and Soft Matter, Schrödinger
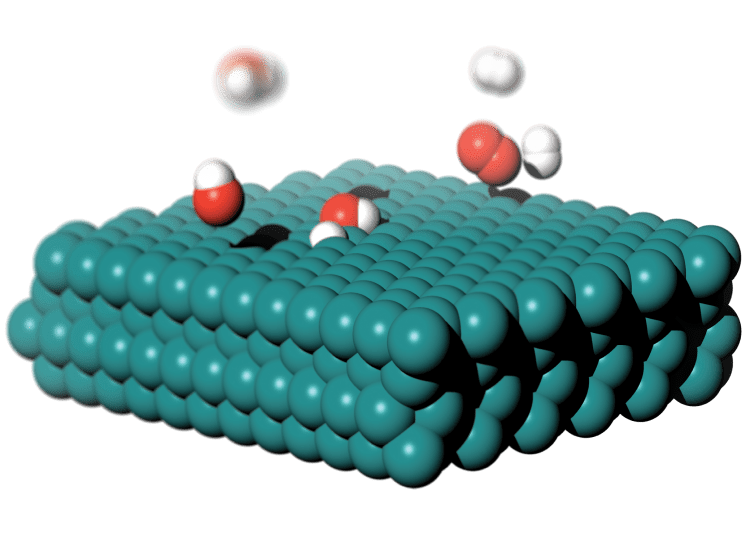
リチウムイオン電池における負極ー電解質界面の解析
シュレーディンガー株式会社 シニア サイエンティスト 井本 文裕
閉会
懇親会
※海外からの講演者は英語での発表となります。
※Presentations by Japanese speakers are available in Japanese only.
【開催形式と会場】
・現地開催です。オンライン配信はございません。
会場
鉃鋼エグゼクティブラウンジ&カンファレンスルーム
〒100-0005 東京都千代田区丸の内1丁目8-2, 鉃鋼ビルディング 南館, 4階
アクセス
※会場参加の登録受付は9月25日(水)23:59までといたします。
※会場の収容可能人数には限りがあり、登録受付期日前であっても、上限に達し次第締め切りとなります。お早めにお申し込みください。
※会場参加者様へは、別途メールにて詳細をご案内いたします。
【参加費】
無料
【お申込み方法】
▼参加のお申し込みはこちらから▼
https://form.run/@schrodinger-20241003
所属企業または所属機関のメールアドレスにて、ご登録をお願いします。
所属が明らかでない、また、個⼈メールアドレスでご登録の場合などは、出席をご遠慮いただく場合がございますのであらかじめご了承ください。参加お⼀⼈様につき⼀登録をお願いします。
同業他社さまには参加をご遠慮頂いております。申し訳ございませんが、ご理解のほど宜しくお願い致します。
※ご質問、ご不明な点がございましたら下記までお問い合わせください。シュレーディンガー株式会社 UGM事務局
E-mail: info-japan@schrodinger.com