Release Notes
Release 2023-2
Small Molecule Drug Discovery
Platform Environment
Maestro Graphical Interface
- Added support for interactive nonstandard protein residue mutation via the Workspace context menu [2023-2]
- Save up to 5K resolution GIFs (Workspace -> Animate) [2023-2]
- Reduce unexpected visual clutter with new preference to limit the number of atom labels [2023-2]
- Maestro to PyMOL connection [2023-2]
- Create a simple PyMOL movie from Maestro scenes
- Maestro to LiveDesign connection [2023-2]
- Connect to LiveDesign servers with new access point and connection status indicator
- Retain and reuse single sign on (SSO) tokens to streamline the login process
- Benefit from a streamlined process of pushing Maestro data into LiveDesign via the Export panel
- Export Structures – New option to export MD-ready structure / trajectory to CMS file [2023-2]
- New “Show Atom Properties (beta)” panel [2023-2]
- List multiple atom properties for selected atoms
- Display SMARTS/SMILES patterns for contiguous selected atoms, including the SMARTS index of the hovered-over atom
Workflows & Pipelining [KNIME Extensions]
- Includes the latest version of KNIME [2023-2]
- Improved usability of the Extract Properties node configuration panel [2023-2]
- Updated LiveDesign nodes and protocols: [2023-2]
- Adaptable input column checking of the LiveDesign input node
- KNIME protocol section to install extra KNIME extensions is more robust
Target Validation & Structure Enablement
Protein Preparation
- Bond orders and charges can now either be re-assigned or only added to ligands and residues with missing bond orders [2023-2]
Protein X-Ray Refinement
- Improved robustness when running Phenix/OPLS [2023-2]
- New plots to inspect the best refinement statistics from Phenix/OPLS weight scans [2023-2]
- Phenix/OPLS weight scan automatically choses a near-optimal combination of refinement parameters [2023-2]
AlphaFold Download / Process
- Downloaded AlphaFold structures are returned with an automatically created heatmap of the PAE matrix [2023-2]
- Processed AlphaFold structures can now be optionally capped and the pLDDT threshold changed [2023-2]
Multiple Sequence Viewer/Editor
- New ability for the detection and annotation of Vernier zone residues in antibodies [2023-2]
IFD-MD
- Added visual indicator when the target ligand is missing torsional parameters [2023-2]
Desmond Molecular Dynamics
Improved plotting for Trajectory Plots [2023-2]
Hit Identification & Virtual Screening
Ligand Preparation and Conformation Generation
- Full support for Epik 7 pKa predictions within the LigPrep interface and command line invocation [2023-2]
Empirical and QM-based pKa Prediction
- Conjugate acid/base labels are now included in the Epik 7 log file [2023-2]
Hit Analysis
- Release of the new Hit Analysis interface to streamline interactive analysis and selection of hits from virtual screening campaigns based on molecular properties, ligand feature locations, and shape alignments [2023-2]
Active Learning Applications
- Added ability to train on preexisting FEP data for relative and absolute AL-FEP [2023-2]
- Added histogram of compounds prioritized by machine learning for relative and absolute AL-FEP [2023-2]
- Modified default machine learning settings to improve out-of-the-box performance of relative and absolute AL-FEP [2023-2]
ABFEP
- New capability for fast filtering of inactive ligands to dramatically improve ABFEP throughput [2023-2]
Lead Optimization
Macrocycles
- Improved atom mapping in FEP+ maps for macrocycles [2023-2]
FEP+
- New functionallity to auto-populate state populations in the FEP+ interface [2023-2]
- Improved reporting of results in a single column (val ± err) in the overview tab of the FEP+ interface [2023-2]
- Option to automatically merge force field parameters generated during an FEP+ job with those defined in your Maestro preferences [2023-2]
- Option to select alternative water models for Relative Binding FEP+ via Advanced settings [2023-2]
- Fast filtering of inactive ligands to dramatically improve ABFEP throughput [2023-2]
- Improved atom mapping in FEP+ maps for macrocycles [2023-2]
Protein FEP
- Ability to simultaneously predict protein thermostability with every protein/ligand selectivity simulation [2023-2]
- Option to interactively edit perturbation topologies for protein residue mutations in FEP+ interface [2023-2]
Constant pH Simulations (Beta)
- Greatly improved accuracy in protein pKa predictions through improved sampling of the physical end states of titratable residue side chains in FEP+ [2023-2]
Quantum Mechanics
- New ability to plot excited state energies in rigid and relaxed coordinate scans [2023-2]
- Support for alignment based on uniform scaling in the VCD/IR spectrum_align utility [2023-2]
Semi-Empirical Quantum Mechanics
- Capability to use GFN2-xTB from within Jaguar and in Jaguar-based workflows [2023-2]
Biologics Drug Discovery
- Improved protein descriptor calculation throughput with ability to run in parallel over multiple CPUs [2023-2]
- Up to 5x speedup in protein surface calculations [2023-2]
- Modeling of single-chain Fvs is now incorporated into the antibody structure-prediction interface [2023-2]
- Modeling of F(ab)2 formats is now integrated into the antibody structure-prediction interface [2023-2]
- MSV is now accessible directly from the protein-protein docking interface [2023-2]
- Selected entries in antibody database management interface can now be exported to MSV and Maestro [2023-2]
Materials Science
GUI for Quantum ESPRESSO
Product: Quantum ESPRESSO (QE) Interface
- Quantum ESPRESSO: Reduced disk usage with hybrid functionals [2023-2]
- Quantum ESPRESSO: Option to import only the final structure in QE import GUI [2023-2]
- Quantum ESPRESSO: Automatic q-point mesh setup with hybrid functional [2023-2]
Transport Calculations via MD simulations
Product: MS Transport
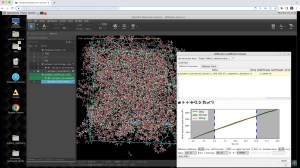
- Diffusion Coefficient Viewer: Visualization of atoms selected for diffusion tracing [2023-2]
- Viscosity: Expanded range of shear stress available for analysis [2023-2]
Materials Informatics
Product: MS Informatics
- Machine Learning Property: Report of entries with failed predictions if any [2023-2]
- Machine Learning Property: Density prediction for molecular liquids [2023-2]
- Machine Learning Property: Models to measure uncertainties from the predicted properties [2023-2]
Coarse-Grained (CG) Molecular Dynamics
Product: MS CG
- CG FF Builder: Better defaults for convergence [2023-2]
- CG FF Builder: Option to set initial values [2023-2]
- CG FF Builder: Option to save the force field file in viewer [2023-2]
Dielectric Properties
Product: MS Dielectric
- Complex Permittivity: Option to adjust the length of dipole moment sample extracted from the source trajectory (command line) [2023-2]
MS Maestro Builders and Tools
- Crystal: Edit option for lattice parameters when importing PDB without them [2023-2]
- Manipulate Cell: Option to take multiple entries as input for selected operations [2023-2]
- Meta Workflows: Support for molecular QM simulation stages [2023-2]
- Meta Workflows: Option to compute ESP charges for subsequent stages [2023-2]
- Semicrystalline Polymer: Improved robustness and speed for building semicrystal interface [2023-2]
Classical Mechanics
- Crystal Morphology: Improved pop-up guideline for setting a proper input cell size [2023-2]
- Improved UI to select atoms for substrate restraints in MD-based workflow panels [2023-2]
- Droplet: Option to take existing MD simulation trajectory as input [2023-2]
- Droplet: Support for built-in and custom solvents for contact angle prediction [2023-2]
- Electroporation: Workflow module to simulate and assess membrane electroporation [2023-2]
- Evaporation: Redesigned UI for the workflow setup panel [2023-2]
- Evaporation: Support for evaporating multiple solvents [2023-2]
- Evaporation: Added flexibility to evaporation zone definition [2023-2]
- Evaporation Results: Load structures from one or more iterations into the Project Table [2023-2]
- MD Multistage: Support for negative external electric field [2023-2]
- MD Multistage: Option to remove center of mass velocity [2023-2]
- Stress Strain: Support for sinusoidal loading (command line) [2023-2]
Quantum Mechanics
- Adsorption Enumeration: Option to set bridging and hollow sites for adsorption [2023-2]
- Complex Enumeration: Report of the ligand exchange stability in Project Table [2023-2]
- Prediction of singlet excitation energy transfer rates (SEET) (command line) [2023-2]
Education Content
- New Tutorial: Calculating Voltage Curves on Spinel Intercalation Compounds [2023-2]
- New Tutorial: Machine Learning for Ionic Conductivity [2023-2]
- New Tutorial: Electroporation [2023-2]
- Update: Evaporation [2023-2]
- Update: Droplet Contact Analysis [2023-2]
- Update: Viscosity [2023-2]
- Update: Machine Learning Property Prediction [2023-2]
LiveDesign
What’s New in 2023-2
Kubernetes versions of LiveDesign include:
- Composite Rows
- Create relationships between entities to better view the composition of complex mixtures, linking them as subcomponents and showing them as indented rows.
- Get a better understanding of drug formulations by viewing the components that make up the mixture
- Better define the composition of stereoisomeric mixtures
- Analyze the differences between different battery electrolyte formulations
All versions of LiveDesign include:
- Forms
- Show data in a custom, dense arrangement with the Matrix widget
- Search for compound IDs directly in the Compound Image widget
- Set up forms more quickly, and identify columns to add to widgets more quickly, with an updated column shuttle
- Collapse or expand all swimlanes in Kanban widgets using a menu option
- Form widget titles automatically expand to show longer widget titles when a single widget is within a window
- View any custom Experimental Metadata in the assay tooltips
- Any metadata can be added to LiveDesign from corporate assay capture systems through the Data Integrator
- Generic Entity – store, model, and analyze any kind of modality in LiveDesign
- Purge and Overwrite experimental data
- Append new data to a Lot
- 3D Visualizer
- View halogen bond and salt bridge interactions
- Show Chain ID in the residue label
- Significant improvements to streamlining integration of Maestro sessions and LiveDesign servers
- Benefit from visual notice in Maestro of connected LiveDesign servers, user account recall, and automatic connection to LiveDesign with valid single sign-on
- UI and UX Improvements
- Independently size column groups and column headers in the LiveReport spreadsheet view
- View column metadata in tooltip by hovering over a column title in the spreadsheet
- View the true data point color in Plots after selecting data points
- Expand and collapse the plot legend to avoid obscuring data points
- Remove compound images from Plot tooltips
- Resize columns in the Assay Data Viewer tool
- Switch between row-per modes more easily
- Use angle bracket and ampersand characters in formulas for manipulating strings, such as the split() formula
- Importing compounds through a file and matching by IDs will now skip rows that do not have a match, and report which IDs failed to import
What’s Been Fixed
- Cell coloring rules no longer extend beyond the edge of a tile
- Data & Columns tree tooltips show the correct “View” button or “Edit” button for columns, based on each users’ assigned permissions
- Date and Datetime display formats set within the Admin Panel now apply to all users with a role set to ‘User’
- Editing a picklist Freeform column value in a Kanban widget will immediately update the tile’s location within the Kanban widget
- Filter conditions for formula columns that include substructure images will correctly show the substructure images
- Filter conditions on columns that have file attachments no longer show file IDs in the suggestion dropdown
- Forms correctly show a pointer cursor instead of a grab cursor when viewing the form
- Forms now support drilldown from Kanban widgets to Spreadsheet and Table widgets
- Forms now support multiple instances of the same custom tool
- Histogram and Pie plots permit creating a defined number of equally distributed bins
- Histogram plots within Forms now permit defining custom bins
- LiveDesign will start even if the preprocessor config includes unsupported fields
- MPO desirability cell borders no longer show a color when the color is defined by a proxy value
- MPO tooltips now appear in Form widgets that have a drilldown selection
- Picklist Freeform columns now permit bulk copying dates from assay columns
- Plots that use the Highlighted Substructure column will now show compound images when defining custom bins in Histogram and Pie charts
- Plots with a regression line will correctly scale when the plot axes are converted to log scale
- Plots with log axes no longer show negative values
- Reagents with numeric IDs will correctly carry through their data when used within Reaction Enumeration
- Scatter plots with three axes and many data points no longer show blank exports
- Selecting a range of tiles in a Kanban widget, while holding down the shift key on the keyboard, will now only select the visible tiles within a vertical
- The maximum number of data points allowed within a plot, set within the Admin Panel, now applies to all users with a role set to ‘User’
- Toggling to different plot tabs in Forms will now show the correct data point tooltips
- Typed text within filter conditions, that has not been saved, is now removed after selecting an option from the dropdown list
Training & Resources
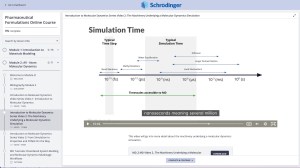
Online Certification Courses
Level up your skill set with hands-on, online molecular modeling courses. These self-paced courses cover a range of scientific topics and include access to Schrödinger software and support.
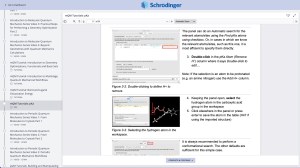
Tutorials
Learn how to deploy the technology and best practices of Schrödinger software for your project success. Find training resources, tutorials, quick start guides, videos, and more.





























Trusted by global leaders in industry